Optimization of Parameters for the Production of Cellulase from Achromobacter xylosoxidans BSS4 by Solid-State Fermentation
Erandapurathukadumana Sreedharan Hareesh, Panichikkal Abdul Faisal, Sailas Benjamin
Erandapurathukadumana Sreedharan Hareesh, Panichikkal Abdul Faisal, Sailas Benjamin*
Enzyme Technology Laboratory, Biotechnology Division, Department of Botany, University of Calicut, Kerala, India
Received date: August 17, 2016; Accepted date: August 31, 2016; Published date: September 07, 2016
Citation: Hareesh ES, Faisal PA, Benjamin S. Optimization of Parameters for the Production of Cellulase from Achromobacter xylosoxidans BSS4 by Solid-State Fermentation. Electronic J Biol, 12:4.
Abstract
Now-a-days, an increase in demand for the costeffective methods in the production of industrially useful enzymes like cellulases is witnessed. Thus, the present study was aimed at optimizing the growth parameters like temperature, pH, moisture and duration of incubation for the production of cellulase by Achromobacter xylosoxidans strain BSS4, employing solid-state fermentation (SSF) strategy. Substrates like rice bran, corn bran, sugarcane bagasse, rice straw and saw dust were selected for examining the most efficient one among them in supporting the maximum cellulase production. From the substrates, corn bran supported the maximum cellulase activity (107.7 U/gds) under unoptimized condition; hence, corn bran was selected as substrate for the statistical optimization of different parameters for maximizing the cellulase yield. Box-Behnken design coupled with response surface methodology was applied for the statistical optimization of the process parameters [temperature (40°C), pH (5), moisture (60%), incubation time (6 h)]; which resulted in a 3.7 fold increase in cellulase activity, i.e., from 107.7 U/gds, the activity increased to 512.98 U/gds. Briefly, this study shows that A. xylosoxidans BSS4 is a good producer of cellulase under SSF using corn bran as the substrate, which paves a way for the exploitation of A. xylosoxidans strain BSS4 with an industrial perspective.
Keywords
Achromobacter xylosoxidans BSS4; Cellulose; Solid-state fermentation; Box-Behnken design; Response surface methodology.
Introduction
Production of waste materials is an undisputable part of human behaviour; which is emerged from several domains including industries, forestry, agriculture, etc. Lignocellulose constitutes the core components of different waste streams, and is being effectively used for the production of biofuels, biogas, enzymes, etc. [1]. Cellulose is a cell wall polysaccharide which continually replenished in nature by photosynthesis, gone waste in the form of agricultural losses, and wastes of food processing industry, etc. [2]. Cellulose is a linear polysaccharide of glucose residues connected by β-1,4, glucosidic linkages [3]. Degradation of cellulosic biomass is performed by a mixture of hydrolytic enzymes, collectively known as cellulases [4]. Complex cellulase enzyme systems are classified in to glycoside hydrolase families, and can be divided in to three types: (a) endo-1,4- β-D-glucanase or endoglucanase (EC.3.2.1.4), (b) 1,4-β-D-cellobiohydrolase or exoglucanase (EC.3.2.1.9) and (c) β-glycosidase (EC.3.2.1.21); they in unison pave the way to degrade the cellulosic substrates effectively [5]. Microorganisms play the pivotal role in assimilating the cellulosic waste. Species of several mycelial fungal genera like Humicola, Aspergillus and Penicillium are the best known producers of cellulase; in addition to several bacterial species belong to genera like Achromobacter, Bacillus, Clostridium, Pseudomonas, Lactobacillus, Serratia and Streptococcus [3]. Bacterial cellulases have several advantages over fungal cellulases, especially their activity and stability under extreme conditions [6].
Solid-state fermentation (SSF) is a much relevant, cheapest and eco-friendly fermentation strategy compared to the submerged fermentation (SmF). In SSF, the substrates are solid and insoluble in nature, e.g. corn fibre, corn stover, sugarcane bagasse, rice hulls, woody crops, etc. [7]. In addition to providing nutrients, solid support and ample aeration to the growing microbes, less water requirement and higher product yield are the unique advantages of SSF; while SmF requires soluble feed stocks like glucose and hence it requires more water and less substrate requirements [7].
Employment of statistical tools for optimizing process parameters has been in use for several decades. The classical method of optimization generally deals with the one at a time strategy, which does not address the combinatorial effect of parameters, and thus leads to the misinterpretation of results [8]. These days, modern statistical optimization system is being used to overcome this problem. Response Surface Methodology (RSM) is considered as the most promising tool in the optimization of process parameters for better yield. RSM is a collection of mathematical and statistical techniques used for analyzing and modelling the influential parameters on to the response of interest such as nutrient requirements and enzyme yield. Plackett-Burman Design (PBD), Central Composite Design (CCD), and Box-Behnken Design (BBD) are the major tools adopted in RSM modelling. RSM is selected in this study as a tool to obtain the most suitable process conditions which control the growth of the microorganism and the enzyme yield. Considering all these facts in concern, the present work focuses on the statistical optimization of growth parameters for cellulase production by Achromobacter xylosoxidans strain BSS4, employing SSF strategy.
Materials and Methods
Chemicals
Analytical and bacteriological grade chemicals from Merk (India) and HiMedia (India) were used for the preparation of the medium for the bacterial culture, and reagents for enzyme assay.
Microorganism
Achromobacter xylosoxidans strain BSS4 (Genbank Accesion No. JQ 407052) described from the Enzyme Technology Laboratory, Department of Botany, University of Calicut, Malappuram, Kerala was used for the present study. Stock cultures were maintained on agar slants prepared in the basal salt medium (BSM), supplemented with carboxymethyl cellulose (CMC) as carbon source [9]. The BSM contained (g/L): 2.0 NaNO3; 1.0 K2HPO4; 0.5 KCL; 0.5 MgSO4.7H2O; 2.0 proteose peptone; 20 agar and 0.5% CMC.
Cellulase production by SSF
Cellulose containing substrates like Corn Bran (CB), Rice Bran (RB), Rice Straw (RS), Sugarcane Bagasse (SB), and Saw Dust (SD) were collected from the local market for the production of cellulase by SSF. Sun-dried and finely ground substrates were kept in the air tight plastic containers to avoid contamination. Fermentation was carried out in conical flask (100 mL) containing 5 g of selected substrate and 10 mL of BSM (substrate to moisture ratio was 1:2). All flasks with required preparations were autoclaved (121°C) for 15 min and inoculated with 0.1mL of inoculum (about 107 cfu) under aseptic condition. After inoculation, all the flasks with media and inoculum or control were incubated at 37°C under static condition. Production of cellulase was assayed at regular intervals of 6 h for 36 h. Before measuring cellulase activity, 5 g of the substrate (from whole flask) was taken to calculate the dry weight. All the experiments were carried out in triplicates.
Crude enzyme extraction
After required incubation, the fermented matter in the culture flasks was used for cellulase assay at regular intervals of 6 h. For the extraction of crude cellulase, 10 mL of 0.1 M citrate buffer (pH-4.8) was added to 5 g fermented matter, and stirred for 10 min. Then the mixture was centrifuged at 6000 × g at 4°C. The clear supernatant (crude enzyme) so obtained was used for cellulase assay.
Cellulase assay
Cellulase was assayed by employing the 3,5-dinitrosalicylic acid (DNS) method of Miller [10]. The reaction mixture contained 0.5 mL of 1% CMC (1 g CMC in 100 mL of 0.1 M citrate buffer, pH 4.8) as substrate; 0.05 mL of crude enzyme (supernatant) and 0.45 mL citrate buffer were added to it and incubated for 30 min in a water bath at 50°C. Subsequently, 3 mL of DNS was added and incubated for 5 min in the boiling water bath for colour development and cooled rapidly. The activity was measured against a reagent blank at 540 nm in a UV-Vis spectrophotometer (Shimadzu, Japan). One unit of cellulase activity is defined as the quantity of cellulase required to liberate 1 μmol of glucose equivalents per minute under the assay conditions. Cellulase activity was calculated using the following formula.
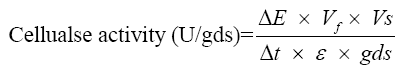 (1)
(1)
ΔE: Absorbance at 540 nm; Vf: Final volume including DNS; Vs: Volume of cellulase used; Δt: Time of hydrolysis; ε: Extinction co-efficient; gds: Dry weight in grams of the substrates.
Statistical optimization of cellulase production
The BBD was used to predict the combined effect of different independent parameters such as temperature, pH, moisture and incubation time on the production of cellulase. The software Minitab version 14 (Minitab USA) was used for designing and analyzing the experimental trials of BBD.
Four culture parameters such as temperature (28, 34 and 40°C), pH (5, 7 and 9), moisture (20, 40 and 60%), and incubation time (6, 12 and 18 h) were selected for BBD. The analysis was made at three levels (high, medium, low), represented by +1, 0 and -1, respectively. A set of 27 experimental trials were suggested by the Minitab. All the trials were carried out in duplicates and the results were analyzed by fitting to a second order polynomial equation (2). Each experimental trial was set up and cellulase was harvested at proper intervals to measure the cellulase activity as per the design.
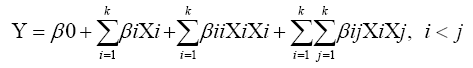 (2)
(2)
Where, Y represents the response variable, β0 is the interception coefficient, βi is the coefficient of the linear effect , βii is the coefficient of quadratic effect, βij is the coefficient of interaction effect when i<j and k is the number of variables.
Validation experiments
To check the validity of quadratic model, 4 experiments as predicted by point prediction software Minitab 14 were performed. Cellulase activity was estimated and compared with the predicted values.
Results
Effect of different substrates on cellulase activity
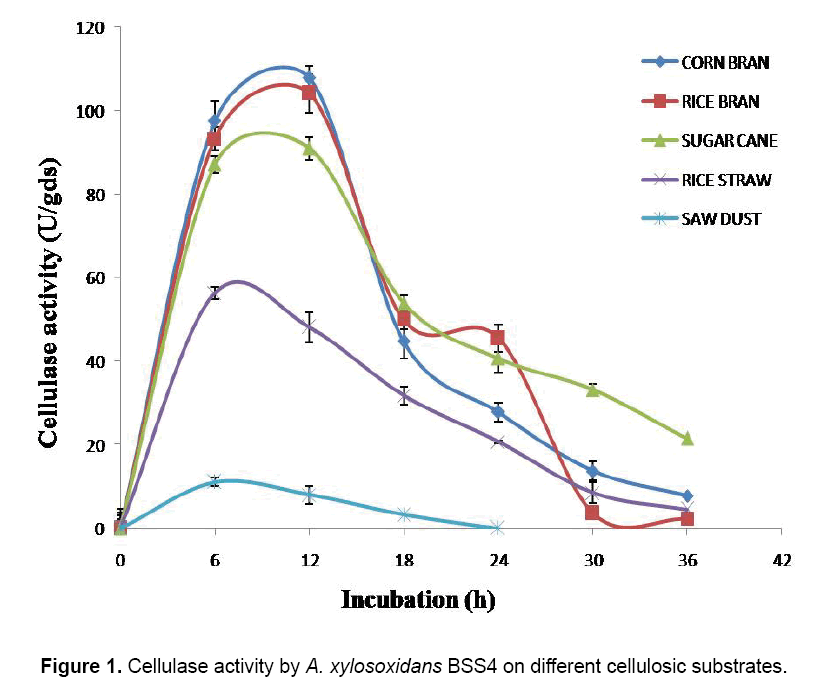
Five different natural substrates (CB, RB, RS, SB and SD) were used for assessing their efficacy as solid support and nutrient source in the production of cellulase (Figure 1). With CB as a substrate, the maximum cellulase activity (107.77 U/gds) was at 12 h of incubation; after that a gradual decrease was observed; while with RB as a substrate, the maximum cellulase activity (104.15 U/gds) was noticed at 12 h of incubation, subsequently the activity decreased gradually; and at 36 h of incubation, the cellulase activity was only 2.37 U/gds. RS supported the maximum production of cellulase at 6 h of incubation (56.42 U/gds), then the cellulase activity was decreased gradually. In the case of SB, the maximum activity (91.1 U/gds) showed at 12 h of incubation, then a gradual decrease occurred; and at 36 h, the activity was 21.4 U/gds; the maximum (11.09 U/gds) activity of cellulase with SD was observed at 6 h of incubation, followed by a sudden fall. Effects of CB (107.77 U/gds), RB (104.15 U/gds) and SB (91.1 U/ gds) were comparable to each other, but the other two substrates (RS and SD) showed much lesser activity, i.e., 56.42 U/gds and 11.09 U/gds, respectively. Of these substrates, CB supported the maximum cellulase activity (107.77 U/gds) (Figure 1).
Statistical optimization of parameters for cellulase production
From the results of screening (Figure 1), CB was selected the best substrate for the optimization process. Four parameters (temperature, pH, moisture and incubation) were subsequently considered for BBD analysis followed by RSM to estimate the optimum combination of these parameters for maximizing the production of cellulase. A set of 27 experiments was conducted according BBD and the results showed that the predicted and the experimental values for cellulase activities were not significantly different (Table 1); i.e., the R2 value was 0.98 - close to unity. A second order polynomial equation was fitted to the cellulase activity, which resulted in the following regression Equation.
| Run order | Temperature (°C) | pH | Moisture (%) | Incubation time(h) | Observed (U/gds) | Predicted (U/gds) |
|---|---|---|---|---|---|---|
| 1 | 28 | 5 | 40 | 12 | 260.61 | 258.6 |
| 2 | 40 | 5 | 40 | 12 | 251.86 | 267.309 |
| 3 | 28 | 9 | 40 | 12 | 193.06 | 168.311 |
| 4 | 40 | 9 | 40 | 12 | 187.39 | 180.1 |
| 5 | 34 | 7 | 20 | 6 | 36.22 | 74.73 |
| 6 | 34 | 7 | 60 | 6 | 413.42 | 436.162 |
| 7 | 34 | 7 | 20 | 18 | 21.93 | -10.109 |
| 8 | 34 | 7 | 60 | 18 | 457.29 | 409.48 |
| 9 | 28 | 7 | 20 | 12 | 39.46 | 59.188 |
| 10 | 40 | 7 | 20 | 12 | 18.1 | 3.322 |
| 11 | 28 | 7 | 60 | 12 | 354.47 | 383.583 |
| 12 | 40 | 7 | 60 | 12 | 465.34 | 459.948 |
| 13 | 34 | 5 | 40 | 6 | 309.48 | 286.99 |
| 14 | 34 | 9 | 40 | 6 | 216.71 | 208.041 |
| 15 | 34 | 5 | 40 | 18 | 218.02 | 241.03 |
| 16 | 34 | 9 | 40 | 18 | 105.65 | 142.48 |
| 17 | 28 | 7 | 40 | 6 | 272.74 | 249.188 |
| 18 | 40 | 7 | 40 | 6 | 260.21 | 253.699 |
| 19 | 28 | 7 | 40 | 18 | 186.19 | 187.689 |
| 20 | 40 | 7 | 40 | 18 | 185.13 | 203.676 |
| 21 | 34 | 5 | 20 | 12 | 39.85 | 29.697 |
| 22 | 34 | 9 | 20 | 12 | 26.20 | 24.958 |
| 23 | 34 | 5 | 60 | 12 | 507.99 | 504.218 |
| 24 | 34 | 9 | 60 | 12 | 326.31 | 331.458 |
| 25 | 34 | 7 | 40 | 12 | 211.83 | 226.152 |
| 26 | 34 | 7 | 40 | 12 | 233.53 | 226.152 |
| 27 | 34 | 7 | 40 | 12 | 233.08 | 226.152 |
Table 1: Experimental trials designed according to Box-Behnken model for the optimization of cellulase production.
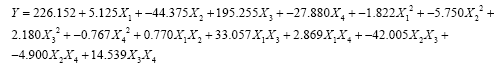 (3)
(3)
Where X1-temperature, X2-pH, X3-moisture, X4- incubation time.
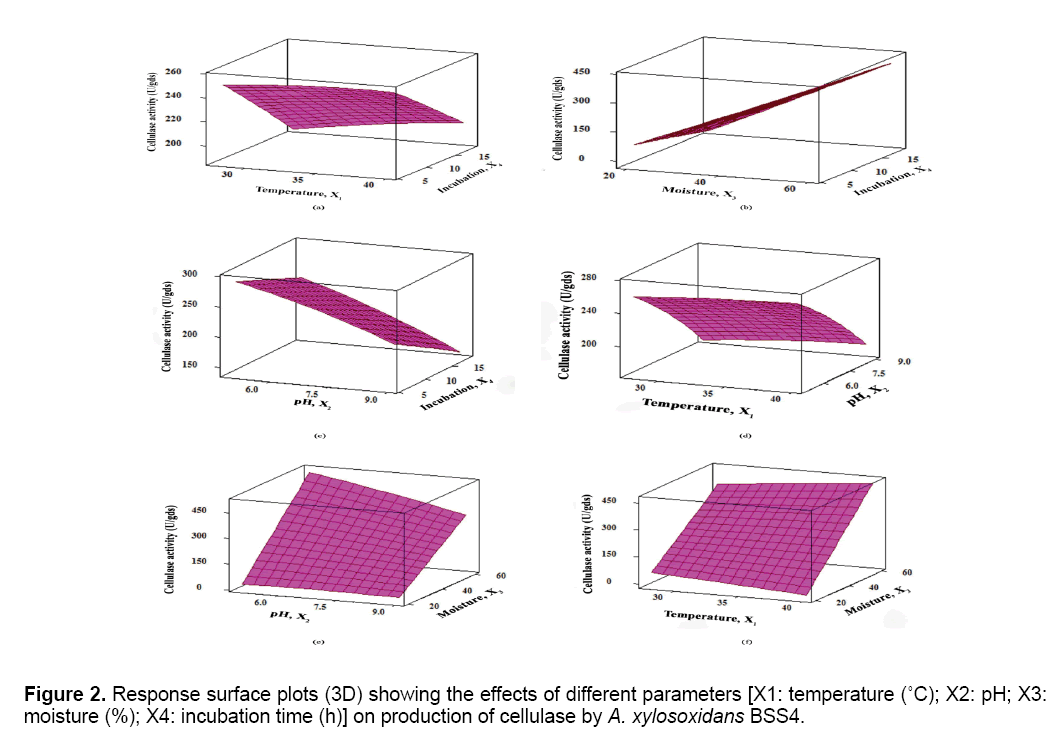
The results were analyzed by ANOVA (Table 2). Based on these results, the model was utilized to generate response surfaces for the analysis of the variable effects on the production of cellulase. The response surface plots obtained as per the Equation (3) are depicted in Figure 2.
| Source | DF | Seq SS | Adj SS | Adj MS | F | P |
|---|---|---|---|---|---|---|
| Regression | 14 | 503457 | 503457 | 35961 | 37.30 | 0.000 |
| Linear | 4 | 490768 | 490768 | 122692 | 121.27 | 0.000 |
| Square | 4 | 283 | 283 | 71 | 0.07 | 0.989 |
| Interaction | 6 | 12406 | 12406 | 2068 | 2.14 | 0.123 |
| Residual Error | 12 | 11568 | 11568 | 964 | ||
| Lack-of-Fit | 10 | 11261 | 11261 | 1126 | 7.33 | 0.126 |
| Pure Error | 2 | 307 | 307 | 154 | ||
| Total | 26 | 515025 |
Table 2: Analysis of variance for the second order polynomial model in optimizing cellulase production. (DF: Degree of Freedom;Seq SS: Sequential Sum of Square;Adj SS: Adjacent Sum of Square;Adj MS: Adjacent Mean Square; F: text of statistics; P: Probability)
Validation of cellulase production
Four random experimental conditions were evaluated for the validation of the model. In all these instances, model prediction was in good agreement with the experimental data, and correlation coefficient was found to be 0.99 (Table 3). Correlation coefficient was close to 1.0, suggesting the significance of the model. At this state, the optimum production of cellulase was found to be 512.98 U/gds (at 40°C, pH 5, moisture 60% and incubation period 6 h). Thus, the statistical optimization resulted in the 3.7 fold increase of cellulase activity over the unoptimized condition.
| Sl. No. | Temperature (°C) | pH | Moisture (%) | Incubation time (h) | Observed (U/gds) | Predicted (U/gds) |
|---|---|---|---|---|---|---|
| 1 | 40 | 5 | 60 | 6 | 512.98 | 544.23 |
| 2 | 28 | 6 | 60 | 12 | 435.10 | 424.43 |
| 3 | 28 | 6 | 40 | 10 | 222.54 | 251.55 |
| 4 | 37 | 6 | 40 | 10 | 236.83 | 259.17 |
Table 3: Experimental trials for validation of the predicted model.
Discussion
In the present work, employing A. xylosoxidans BSS4, different natural cellulosic materials; viz., CB, RB, RS, SB and SD were used as substrates for the production of cellulase. Among these substrates, CB supported the maximum cellulase activity under the unoptimized condition. According to Sukumaran et al. [11] production of cellulase is generally induced by cellulosic substrates and is repressed when easily utilizable sugars are available for growth. SB contains higher free sugars than CB. This may be a reason for the lesser cellulase production with SB as substrate, compared to CB; and surging the screening process, it was observed that all the substrates after 12 h of incubation showed a sudden decrease in the cellulase activity. Sreedevi et al. [9] reported that A. xylosoxidans BSS4 showed the maximum cellulase activity (68.37 U/mL) at 6 h of incubation with CMC as substrate using SmF. In the present case, using same A. xylosoxidans BSS4, the maximum cellulase activity was 107.7 U/gds at 12 h of incubation on CB under unoptimized SSF conditions. Upon Comparison, the present work is a clear proof for the upper hand of SSF over SmF in cellulase production. An increased cellulase activity of 512.98 U/gds was observed on corn bran under optimized parameters like temperature (40°C), pH (5), moisture (60%) and incubation (6 h).
To date, many naturally available substrates such as ground nut shell, rhinoceros dung, agricultural wastes, etc. are being used for the production of cellulose [12-14]. From the BBD experimental results, influential and combinational effects of individual parameters to the cellulase production could be observed. It was reported that, physicochemical factors would influence the growth and production of cellulase by microorganisms, which include chemical composition of the agro residues (cellulose, hemicelluloses, lignin, nitrogen and minerals), presence of an activator or an inhibitor in the agro residues, diffusion of the catabolites, and the type of organism used for fermentation [15]. Liu and Yang reported the maximum cellulase production (23.76 IU/g) by Trichoderma korinjii AS3.4262 upon SSF using lignocellulosic waste from the vinegar industry [16]. Gao et al. [17] reported 581 U/gds cellulase activity on corn stover using a thermo acidophilic fungus Aspergillus terreus M11 under SSF. These results are quite comparable with the present work, indicating that the bacterium can also produce cellulase in the range of a fungus employed for the commercial production of cellulase by SSF under statistically optimized conditions. In classical statistical method, one can only get the information on how a single parameter influencing the product (enzyme) yields. By using modern statistical tool like RSM, it is easier to know the combined effect of parameters on to the response [18]. In the present work, the statistical optimization resulted in 3.7 folds increase of cellulase activity using CB over the unoptimized condition. Thus, these results indicate the excellent scope A. xylosoxidans BSS4 for the production of cellulase with a commercial perspective, employing CB as the substrate.
Conclusion
Corn bran was found as a good substrate for the production of cellulase by A. xylosoxidans BSS4 under SSF. Response surface methodology proved to be a practical, powerful and convenient tool for getting optimum fermentation conditions. So this work concludes that A. xylosoxidans BSS4 may be exploited further with an industrial perspectives, especially with a focus on its potentials for the production of lignolytic enzymes.
Acknowledgement
The financial assistance (File No. 108/SPS/2014/ CSTE) from Kerala State Council for Science, Technology and Environment (KSCSTE), Government of Kerala, is gratefully acknowledged.
References
- Van Dyk JS, Gama R, Morrison D, et al. (2013). Food processing waste: Problems, current management and prospects for utilisation of the lignocellulose component through enzyme synergistic degradation. Renewable and Sustainable Energy Reviews.26: 521-531.
- Das H, Singh SK. (2004). Useful by-products from cellulosic wastes of agriculture and food industry- A critical appraisal. Critical Reviews in Food Science and Nutrition.44:77-89.
- Sajith S, Priji P, Sreedevi S, et al. (2016). An overview on fungal cellulases with an industrial perspective.Journal of Nutrition and Food Sciences.6: 461.
- Sajith S, Sreedevi S, Priji P, et al. (2014). Production and partial purification of cellulase from a noval fungus, Aspergillusflavus BS1. Annuals of Microbiology.64: 763-771.
- Brijwani K, Oberoi HS, Vadlani PV. (2010). Production of a cellulolytic enzyme system in mixed culture solid state fermentation of soybean hulls supplemented with wheat bran. Process Biochemistry.45: 120-128.
- Maki M, Leung KT, Qin W. (2009). The prospects of cellulase producing bacteria for the bioconversion of lignocellulosic biomass. International Journal of Biological Sciences.5: 500-516.
- Benjamin S, Pandey A. (1998). MixedÃÆâÃâââ¬ÃâÃÂsolid substrate fermentation. A novel process for enhanced lipase production by Candida rugosa. ActaBiotechnologica.18: 315-324.
- Priji P, Unni KN, Sajith S, et al. (2014). Production, optimization and partial purification of lipase from Pseudomonas sp. strain BUP6, a novel rumen bacterium characterized from malabari goat. Biotechnologyand Applied Biochemistry.62: 71-80.
- Sreedevi S, Sajith S, Benjamin S. (2013). Cellulaseproducing bacteria from the wood-yards on kallai river bank. Advances in Microbiology. 3: 326-332.
- Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry.31: 426-428.
- Sukumaran RK, Singania RR, Pandey A. (2005). Microbial cellulases-production, applications and challenges. Journal of Scientific and Industrial Research.64:832-844.
- Vyas A, Vyas D, Vyas KM. (2005). Production and optimization of cellulases on pretreated groundnut shell by Aspergillusterreus AV49. Journal of Scientific and Industrial Research.64: 281-286.
- Singh S, Moholkar VS, Goyal A. (2013). Isolation, identification and characterization of a cellulolytic Bacillus amyloliquefaciens strain SS35 from rhinoceros dung. ISRN Microbiology. 1-7.
- Sudto A, Punyathiti Y, Pongsilp N. (2008). The use of agricultural wastes as substrates for cell growth and carboxymethylcellulase production by Bacillus subtilis, Escherichia coli and Rhizobium sp. KMITL Science and Technology Journal.8: 84-92.
- Chinn MS, Nokes SE. (2006). Screening of thermophilic anaerobic bacteria for solid substrate cultivation on lignocellulosic substrates. Biotechnology Progress.22: 53-59.
- Liu J, Yang J. (2007). Cellulase production by Trichodermakoningii AS3.4262 in solid state fermentation using lignocellulosic waste from the vinegar industry. Food Technology and Biotechnology.45:420-425.
- Gao J, Weng H, Zhu D, et al. (2008). Production and characterization of cellulolytic enzymes from the thermoacidophilic fungal Aspergillusterrus M11 under solid state cultivation of corn stover. Bioresource Technology.99: 7623-7629.
- Faisal PA, Hareesh ES, Priji P, et al. (2014). Optimization of parameters for the production of lipase from Pseudomonas sp. BUP6 by solid state fermentation. Advances in Enzyme Research. 2: 125.

Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences