Study on Phosphobacteria Dissolving Phosphorus and Gene Clone and Expression of Thermostable Phytase in Bacteria
Lily Liu
1College of Life Science,Nankai University,Tianjin 300074,P. R. China
2Chemistry and Life Science College,Tianjin Normal University,Tianjin 300074,P.R.China
# Supervisor: Professor Wenbo Yang; Approved date: April 2004.
Abstract
The phosphorus dissolving action of phosphobacteria 9320-SD24 cultured in liquid medium was studied by the available phosphorus assay and the whole phosphorus assay. The available phosphorus was increased by about 1985.45 μmol/L in a liquid medium containing powdered rock phosphate. The transformation rate of the available phosphorus from whole phosphorus was 31.12%. On a solid medium containing powdered rock phosphate, it showed that a transparent sphere of dissolving phosphorus was produced by a colony of phosphobacteria 9320- SD24. In this dissertation the kinetic properties of phosphorus dissolving by the phosphobacteria 9320-SD24 is reported using powdered rock phosphate as the reactive material. The samples were treated with H2SO4-H2O2 for the whole phosphorus assay. The available phosphorus (dissolving phosphorus) in the samples was determined by calorimetry. The amount of livingcells was counted in colony-forming units by cultivating on solid medium. The results indicated that the rate of product formation was positively related to the growth rate of phosphorus bacteria 9320-SD24 and its cell density. The kinetics equation of phosphorus dissolving by the phosphobacterial 9320-SD24: dP/dt=α μX, Kinetics equation of the non-available phosphorus (whole phosphorus) transformation: Y=4.54logX+11.23. Strains SD01N, SD01B, SD01D and SD01X which can decompose phytic acid were screened and isolated from 203 bacteria strains. A pair of primers for myo-Inositol-hexaphosphate-phosphohy– olrolase gene was designed. Using this primer pair, a 1.2Kb band (Figure 2) was amplified by PCR from SD01N genomic DNA. Nucleotide sequence analysis revealed the presence of an open reading frame of 1152bp encoding 383 amino acids. This fragment was inserted into expression vector pQE-30 (Figure 2), and the recombinant plasmid was named pQE-30P. The pQE-30P plasmid was transferred into E.coli M15, to make the transformed bacteria strain SDLiuTP01. The expression of myo-Inositolhexaphosphate- phosphohyolrolase gene in recombination strain SDLiuTP01 was detected by SDS-PAGE electrophoresis, at the same time, the products were compared with E.coli M15 containing the empty vector pQE-30. The enzymatic activity of myo-Inositol-hexaphosphate-phosphohyolrolase in recombination strain SDLiuTP01 was demonstrated and it is stable in the range of 25? to 95?. This myo-Inositol-hexaphosphate-phosphohyolrolase belongs to the thermostable phytase family. Its optimum temperature is 65?. It has two optimum pHs are 4.6 and 7.5. The recombinant plasmid pQE-30P was extracted from SDLiuTPO1. The pQE-30P and pTH315 was double digested by the same enzymes and the plasmid pTH315P was obtained by linking the phytase gene to pTH315. The pTH315P was transformed into phosphobacterium 9320-SD24, which makes of the microorganism fertilizer, via electroporation, to become SDLiuTP02. SDLiuTP02 was able to dissolve phytic acid and cropgrowth was enhanced by the metabolites of SDLiuTP02. B is phytase gene from bacteria strain SD01N by PCR; C is a recombination plasmid pMD18-TP with phytase gene. The expression vector pQE-30 and the recombination plasmid pMD18-TP with phytase gene was enzymolysised by the same two enzymes, becoming the new recombination plasmid pQE-30P via linking pQE-30 and phytase gene. The average value of the increased height of corn plants was 20.0mm.The weight of fresh corn plants was increase by 69.5mg per plant .The weight of dry corn plants was increase by 13.5 mg per plant .The average value of the corn plant roots length was increase by 5.0mm.
Abstract
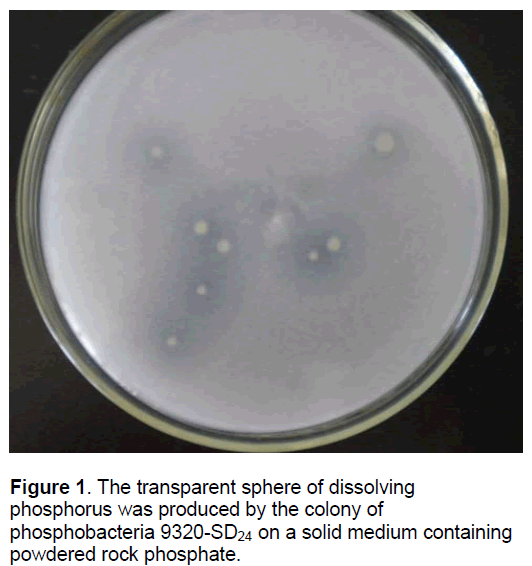
The phosphorus dissolving action of phosphobacteria 9320-SD24 cultured in liquid medium was studied by the available phosphorus assay and the whole phosphorus assay. The available phosphorus was increased by about 1985.45 μmol/L in a liquid medium containing powdered rock phosphate. The transformation rate of the available phosphorus from whole phosphorus was 31.12%. On a solid medium containing powdered rock phosphate,it showed that a transparent sphere of dissolving phosphorus was produced by a colony of phosphobacteria 9320- SD24 (Figure 1).
Keywords
phosphobacteria,phosphorus dissolving kinetics,thermostable phytase,E.coli,clone,powdered rock phosphate,Bacillus sp
In this dissertation the kinetic properties of phosphorus dissolving by the phosphobacteria 9320-SD24 is reported using powdered rock phosphate as the reactive material. The samples were treated with H2SO4-H2O2 for the whole phosphorus assay. The available phosphorus (dissolving phosphorus) in the samples was determined by calorimetry. The amount of living cells was counted in colony-forming units by cultivating on solid medium. The results indicated that the rate of product formation was positively related to the growth rate of phosphorus bacteria 9320-SD24 and its cell density. The kinetics equation of phosphorus dissolving by the phosphobacterial 9320-SD24: dP/dt=α μX,Kinetics equation of the non-available phosphorus (whole phosphorus) transformation: Y=4.54logX+11.23.
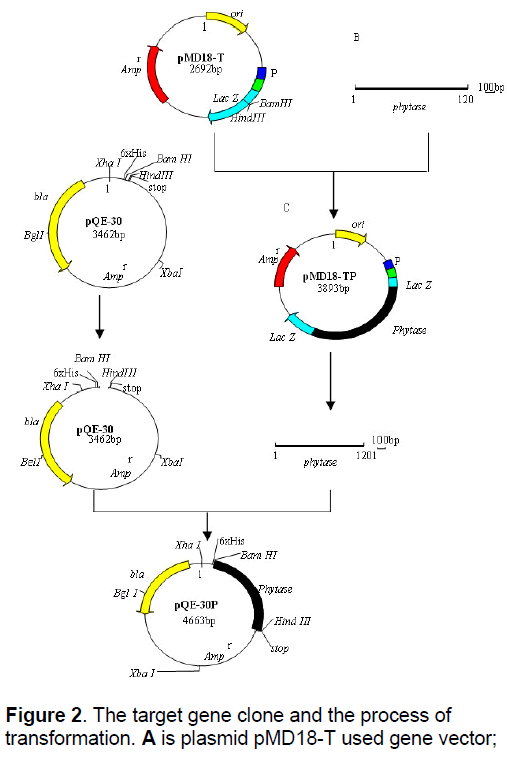
Strains SD01N,SD01B,SD01D and SD01X which can decompose phytic acid were screened and isolated from 203 bacteria strains. A pair of primers for myo-Inositol-hexaphosphate-phosphohy– olrolase gene was designed. Using this primer pair,a 1.2Kb band (Figure 2 ) was amplified by PCR from SD01N genomic DNA.
Nucleotide sequence analysis revealed the presence of an open reading frame of 1152bp encoding 383 amino acids. This fragment was inserted into expression vector pQE-30 (Figure 2 ),and the recombinant plasmid was named pQE-30P. The pQE-30P plasmid was transferred into E.coli M15,to make the transformed bacteria strain SDLiuTP01. The expression of myo-Inositolhexaphosphate- phosphohyolrolase gene in recombination strain SDLiuTP01 was detected by SDS-PAGE electrophoresis,at the same time,the products were compared with E.coli M15 containing the empty vector pQE-30. The enzymatic activity of myo-Inositol-hexaphosphate-phosphohyolrolase in recombination strain SDLiuTP01 was demonstrated and it is stable in the range of 25℃ to 95℃. This myo-Inositol-hexaphosphate-phosphohyolrolase belongs to the thermostable phytase family. Its optimum temperature is 65℃. It has two optimum pHs are 4.6 and 7.5.
The recombinant plasmid pQE-30P was extracted from SDLiuTPO1. The pQE-30P and pTH315 was double digested by the same enzymes and the plasmid pTH315P was obtained by linking the phytase gene to pTH315. The pTH315P was transformed into phosphobacterium 9320-SD24,which makes of the microorganism fertilizer,via electroporation,to become SDLiuTP02. SDLiuTP02 was able to dissolve phytic acid and cropgrowth was enhanced by the metabolites of SDLiuTP02.
The expression vector pQE-30 and the recombination plasmid pMD18-TP with phytase gene was enzymolysised by the same two enzymes,becoming the new recombination plasmid pQE-30P via linking pQE-30 and phytase gene.
The average value of the increased height of corn plants was 20.0mm.The weight of fresh corn plants was increase by 69.5mg per plant .The weight of dry corn plants was increase by 13.5 mg per plant .The average value of the corn plant roots length was increase by 5.0mm.
Acknowledgements
Financial assistance was from grants provided by Tianjin Natural Science Foundation (# 013609811) and Emphasis Item of Science & Technology Project of Tianjin Municipal City in China (# 003122111-4).

Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences