Isolation and Characterization of Acidithiobacillus ferrooxidans Strain FJS from Ramsar, Iran
Samaneh Jahani, Faezeh Fatemi, Mohammad Ali Firoz-e-zare, Mohammad Reza Zolfaghari
1Nuclear Fuel Cycle Research School,NSTRI,Tehran,I.R. Iran
2Department of Microbiology,Faculty of Science,Qom Branch,Islamic Azad University,Qom,Iran
- Corresponding Author:
- Tel: 0989124186349
E-mail: ffatemi@aeoi.org.ir
Received date: September 26,2015; Accepted date: November 02,2015; Published date: November 08,2015
Citation: Jahani S,Fatemi F,Firoz-e-zare MA,et al. Isolation and Characterization of Acidithiobacillus ferrooxidans Strain FJS from Ramsar,Iran. Electronic J Biol,11:4
Abstract
The present work describes a research approach to the aerobic bioleaching of U (IV) ores. Three strains (Acidithiobacillus sp.) isolated from sulfur springs in Ramsar province, Iran and nucleotide sequences of gene fragments of examined Acidithiobacillus sp. strains were recorded in NCBI .Then, uranium bioleaching experiment carried out with native bacteria in different pulp densities. In following, the performance of the isolates was improved progressively by optimizing the pH and initial iron concentrations. The results showed that total uranium extraction was achieved with 100% efficiency using native isolated Acidithiobacillus sp. strain FJ2 in 2.5, 5, 7.5 and 10% pulp densities of the ore. Whereas, the isolates could leach the U (VI) with 68% efficiency at 12.5% ore pulp density. In addition, Acidithiobacillus sp. strain FJ1 leaded to 100% uranium extraction at 2.5% pulp density and 59%at 5% pulp density of the ore. As well, determined optimized values showed that initial ferrous iron concentration at 2.5 g/L and initial pH at 2, gives maximum uranium extraction yields. This is very promising results since, the native isolated strains especially Acidithiobacillus sp. strain FJ2 can be used as a capable bacteria in uranium extraction process.
Keywords
Acidithiobacillus sp.; Isolation; Characterization; Bioleaching; Uranium.
Introduction
Bioleaching,known as biooxidation or biomining,is the use of microorganisms to extract metals from the ores. This process uses a natural ability of microorganisms to transform metals present in the ore in a solid form to a dissolved form [1]. This process is cheaper and easier to conduct in comparison to conventional techniques. Bioleaching of low grade ores is becoming a common alternative since it is an economically viable phenomenon. Moreover,the microorganisms used in these processes are able to grow in highly acidic environments (pH values 1.5 and 3) with high heavy metal ion contents [2]. In addition,the flexibility of microorganisms without problems adapted to different extreme living conditions is a promising advantage [1].
Uranium is an important natural resource used for the generation of nuclear energy,as the raw material [3]. The major minerals of uranium are uraninite,carnotite,pitchblende,coffinite,tobernite,autunite,and tyuyamunite. Extraction of uranium from low grade ore requires large amounts of energy that it is not economical to extract uranium by chemical leaching as the very low content of uranium in weight [4]. So,using of an economic way such as bioleaching for recovery of uranium is very important in industry. Bioleaching of uranium minerals is accomplished by oxidation of the insoluble U4+ form to the acid soluble U6+ form in an acid environment. The indirect mechanism by using ferric ions as an oxidant is proposed for the uranium bioleach process [5]. Meanwhile,some heavy metals such as nickel,aluminum,copper,and zinc extracted in the bioleaching process. Even,those metals can exert a toxic effect if they exceed a certain concentration which again depends on the organism. The toxicity of a metal primarily depends on its interaction with the organism or reactions with biomolecules. Thus,the chemical and physical properties of the metal as well as its concentration can help to predict its toxicity [6].
The microorganisms used for bioleaching are acidophil,such as Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans that gain energy by oxidization of ferrous iron,sulfur and reduced inorganic sulfur compounds [7]. On the basis of the results of 16S rRNA sequence analysis,Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans were combined into a genus,Acidithiobacillus, within the γ-subclass of the Proteobacteria. Various strains of this species have been isolated from natural resources (rocks,ores and mine waters) [8]. It should be noted that biotechnology is an area that has produced a considerable increase in recent years,mainly because occurrence of microbiological techniques and molecular biology,which help to isolate and identify the new native species that are related to more efficient dissolution of minerals [9]. So,in this study,we describe the isolation and identification of native Acidithiobacillus sp. from Iran capable of bioleaching in valuable efficiency. In this way,the effectiveness of the isolated bacteria in uranium extraction was considered at different ore pulp densities (2.5,5,7.5,10 and 12.5%). Also,the performance of the isolates by optimizing the pH and initial iron concentrations were evaluated.
Materials and Method
Sampling sites
The water samples collected from 3 different sulfur springs of Ramsar,Iran (Kash,Khak-e-sefid and Bonyad). The samples were collected from the boiling site of the springs and carried out with sterilized falcons which one-third of space was left blank. Then,the samples were stored in ice box brought to the laboratory.
Isolation and identification of bacteria
Isolation of native Acidithiobacillus was carried out using 9k and Starkey medium [10]. 9k medium contained (gL−1) (NH4)2SO4 3.0,KCl 0.1,K2HPO4 0.5,MgSO4.7H2O 0.5,Ca(NO3)2 0.01,FeSO4.7H2O 44.3. The Starkey medium contained (gL−1) (NH4)2SO4 1.0,CaCl2.2H2O 0.14,KH2PO4 3.0,Mgcl2·6H2O 0.1 and 10 g Sulfur. The pH of the medium was adjusted to 2.0 and 4.0 using 10 N H2SO4 and NaOH,respectively [11,12]. The media were sterilized by an autoclave. Enrichment of microorganisms was carried out with 100 mL liquid medium in 250 mL Erlenmeyer flasks at 30°C on a shaker incubator at 180 rpm. In order to do the purification,solid 9k and Starkey medium with 1.5% agar were used. The plates were incubated for about 1 week at 30°C. By repeating the above plating method,isolated bacteria were obtained. The isolates were identified by phenotypic and genotypic characteristics,which are described below:
Phenotypic characteristics
The bacteria were identified by morphological examinations,biochemical and physiological characterizations. The parameters included colony characteristics,shape,color,motility,Gram’s reaction,oxidase,catalase,urease,H2S and Indole productions,citrate utilization,Methyl-Red test (MR),Voges-Proskauer (V-P) reaction,carbohydrate metabolism (acid-gas production),starch hydrolysis,growth at different pH and temperature,growth in aerobic and anaerobic condition,growth curve and variation of pH and Eh,conducted in accordance with described in Bergey’s Manual of Determinative Bacteria [13].
Genotypic characteristics
Genomic DNA was extracted with DNA extraction kit (Cinnaclone,Iran). In following,16S rRNA were amplified using universal primers (27 F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492 R (5′-GGTTACCTTGTTACGACTT-3′)) [14]. PCR amplifications were carried out using HotStar HiFidelity PCR (Qiagen,Germany). The PCR program consisted of one cycle of DNA denaturation for 5 min at 95°C and then 35 cycles were performed using the following parameters: 1 min at 94°C to denature,1 min at 51.3°C to anneal and 1 min at 72°C to extend,followed by a final extension of 15 min at 72°C. PCR products were analyzed by electrophoresis in 1.5% agarose gel and bands were observed by illuminating Safe stain stained gels on a UV transilluminator. After that,16S rRNA fragments were extracted from agarose gel using gel purification kit (Bioneer,Korea) and PCR products sequenced. Finally,Sequence alignment was performed on the National Center for Biotechnology Information (NCBI) website (https://www.ncbi.nlm.nih.gov/BLAST/) and the phylogenetic trees were constructed.
Mineral characterizations
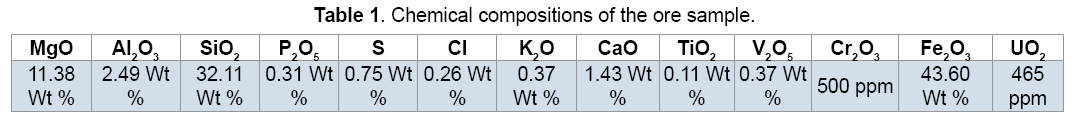
The sample of low grade uranium ore collected from Saghand mining area of Yazd province in Iran. The composition analysis of the ore was carried out by XRF as shown in Table 1. According to the XRF analysis of the ore,main minerals in the ore sample are pyrite,quartz and also uraninite as the main uranium-bearing mineral in the ore.
Uranium bioleaching analysis
Uranium leaching experiments were carried out with the above isolated and characterized bacteria done in 250 mL Erlenmeyer flasks containing 100mL of 9K medium plus different pulp densities (2.5,5,7.5,10,12.5% w/v) of ore powder (d80=170 μm). 10 N H2SO4 was used to adjust the pH value of the leaching solutions. The same numbers of cells were used as inoculums (10%) and the experiments carried out at 30°C fewer than 150 rpm of shaking incubator. 3 mL of samples were removed every day for determining the concentrations of soluble uranium and ferrous iron by an atomic absorption spectrophotometer described by Golmohammadi [15]. When uranium extraction rate reached to 100%,10 mL of the supernatant was removed and added to a 250 mL Erlenmeyer flask containing 90 ml 9k medium with higher pulp density as inoculums. The lost water in the medium was supplemented with 9K medium after each sampling.
Leach liquor compositions
The leach liquor compositions are one of the main factors in uranium extraction. Leach liquor sampling was conducted,when yields of up to 100% uranium extraction were obtained in different pulp densities. Analyses of the leach liquors composition were carried out by inductively coupled plasma-mass spectrometry (ICP-MS) technique (Perkin Elmer).
Optimization of bioleaching process
To achieve the maximum uranium extraction by native efficient bacterium,the composition of medium and growth conditions were optimized by varying two key parameters. The influence of different pH (1.5,2 and 2.5) at 12.5% pulp density was considered into 9K medium in a rotary shaker at 150 rpm and 30°C. In addition,the effect of initial concentrations of Fe2+ (2,2.5 and 3 gL−1) was studied at optimum determined pH. In following,uranium extraction yields were monitored in leach liquors.
Statistical analysis
Data are presented as means Standard Error of Mean (SEM) of three samples (triplicate). The results were subjected to one-way ANOVA followed by Tukey’s HSD using SPSS (version 22) software. Significant levels were defined as P<0.05.
Results
Isolation and identification of the bacteria
In total,six bacteria isolated from different samples of the 9k and Starkey mediums. Between them,3 isolated bacteria were selected based on the best enhanced raise of Eh. These bacteria were considered as efficient sulfur and iron oxidizing bacteria coded as FJ1-3 (FJ1 strain isolated from Kash spring,FJ2 isolated from Khak-e-Sefid spring and FJ3 isolated from Bonyad spring). FJ2 and FJ3 bacteria grew in 9k liquid medium and FJ1 bacterium grew in Starkey liquid medium within 10-12 days. The isolated bacteria spread on 9k and Starkey solid medium. Single colonies appeared on the solid medium after 7 days incubation at 30°C.
Phenotypic characteristics
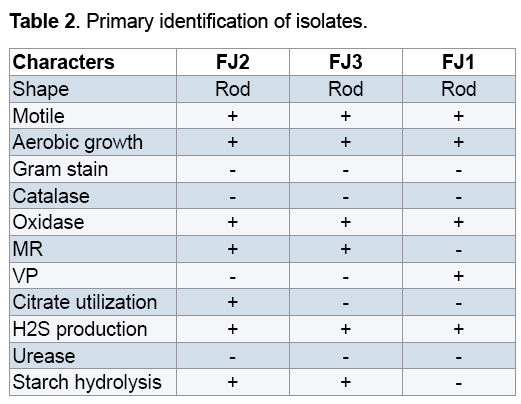
The isolates bacteria subjected to different morphological and biochemical characterization as described in Table 2. FJ2 and FJ3strains colonies on 9k solid medium were round,convex,smooth and brick color. FJ1strain colonies on Starkey solid medium were round,convex,smooth and green in color. Microscopic observations of the isolated bacteria revealed that all of them are rod shaped and motile. The isolates were Gram negative,acidophil,mesophil and aerobic.
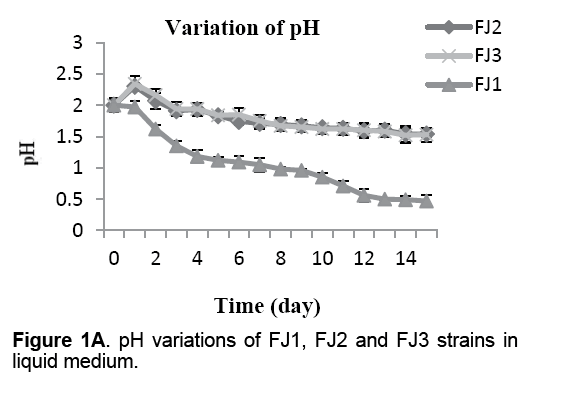
The variations of pH,Eh and growth curve were exhibited in Figures 1A-1C. As it is clear from Figure 1A,amount of pH in 9k medium containing FJ2 and FJ3 strains,increased during the first 24hr of incubation from 2 to 2.4. Afterward,the amount of pH decrease to 1.6 within 15 days. Starkey medium including FJ1 strain didn't have any increase in amount of pH. The initial pH in incubated flask was 2,which was dropped to 0.5 in 15 days.
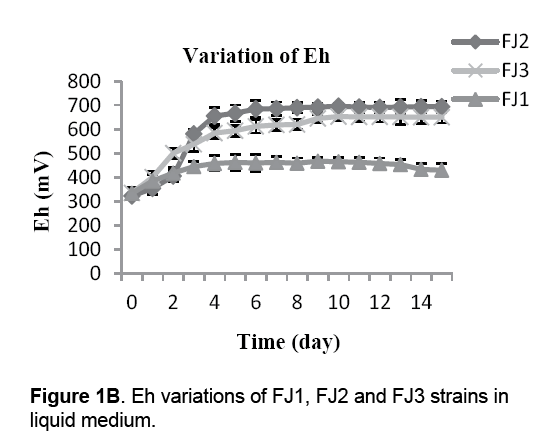
As shown in Figure 1B,Eh value increase was observed in all of the isolated bacteria,triggered by the oxidation of ferrous iron and sulfur. Eh value of 9k medium containing FJ2 strain,begins to rise quickly from 320 mV to 665 mV during 4 days and then rise slowly to 698 mV. While,the Eh value increased from 318 mV to 650 mV (P<0.05) in FJ3 strain. Also,the Eh value of Starkey medium including FJ1 strain increased from 321 mV to 468 mV.
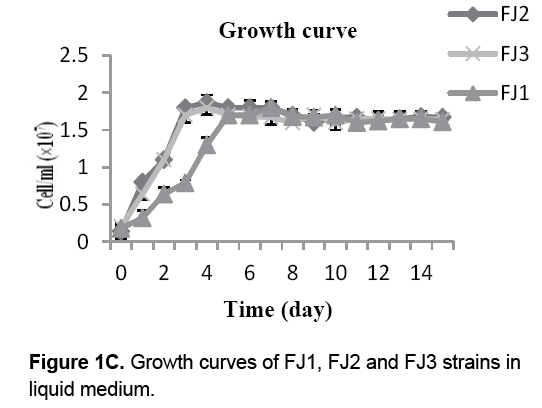
The growth curves for FJ1-3 strains were shown in Figure 1C. The results showed that all three bacteria didn't show any lag phase within 24 hr,whereas after 4 days of incubation,the stationary phase of FJ2 and FJ3 strains were started. Although,after 5 days of FJ1 strain incubated in Starkey medium,the stationary phase was obtained indicating the difference in days of beginning stationary phase. Finally,based on morphological and biochemical tests,the isolated bacteria were identified as Acidithiobacillus sp.
Genotypic characteristics
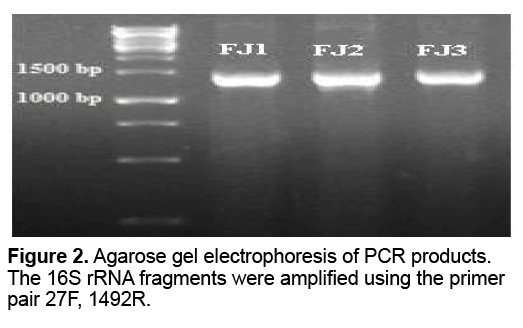
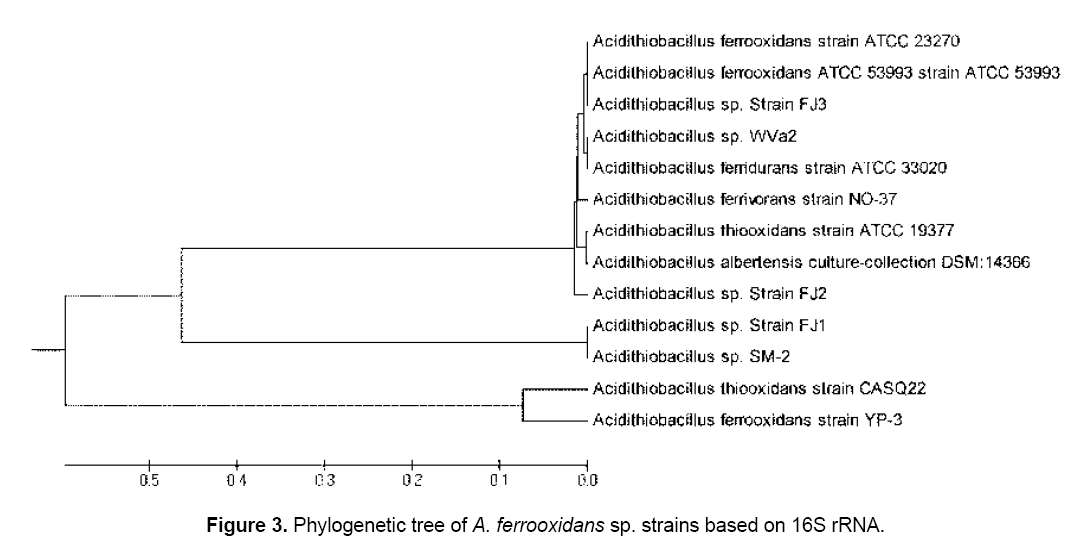
The three isolated were further identified by partial nucleotide sequence of 16S rRNA. Amplification conditions were optimized using genomic DNA from pure cultures of FJ1-3 as templates. Electrophoresis analysis of the PCR products of FJ1-3 showed specific amplification of a single fragment band 1465bp of 16S rRNA (Figure 2 ). The determined partial nucleotides sequences of 16S rRNA were used to find most likeness with the bacterial strains in the GenBank database. A comparative phylogenetic analysis of the 16S rRNA genes of the isolated bacteria were performed with some strains of Acidithiobacillus sp. and theirs phylogenetic tree were shown in Figure 3 . It revealed that the FJ1 strain most closely related to Acidithiobacillus sp. SM-2 with 99% similarity,FJ2 strain was most closely related to Acidithiobacillus ferrooxidans ATCC 23270 with 99% similarity and the FJ3 strain most closely related to Acidithiobacillus ferrooxidans ATCC 53993 with 99% similarity. Nucleotide sequences of gene fragments of examined Acidithiobacillus sp. strains were recorded in NCBI database (accession numbers of deposits KP763724,KP779620 and KP779621).
Uranium bioleaching experiments
The results related to bioleaching experiments (uranium extraction yields,days of extraction and Eh) were indicated in Table 3. Bio-dissolution of uranium was investigated in different pulp densities (2.5- 12.5% (w/v)) at pH 2,35°C with 10% (v/v) inoculums while shaking at 150 rpm. From the results shown in Table 3,it was clear that 100% bio-recovery of uranium were obtained after 2 days incubation at 2.5% (w/v) of pulp density,using FJ2 and FJ3 strains. In addition,at pulp densities of 5,7.5 and 10% (w/v),the 100% of uranium dissolution observed after 5,10,17 days incubation. While,uranium yields was 68% and 60% (P<0.05) at 12.5% pulp density within 20 days,respectively. As shown in Table 3,using FJ1 strain,100% of uranium extraction was observed after 4 days incubation at pulp density of 2.5% (w/v). The amount of uranium extraction at a pulp density of 5% (w/v) was found to be only 59% within 10 days. The results showed that,the rate of bio-recovery of uranium decreased with increasing pulp density.
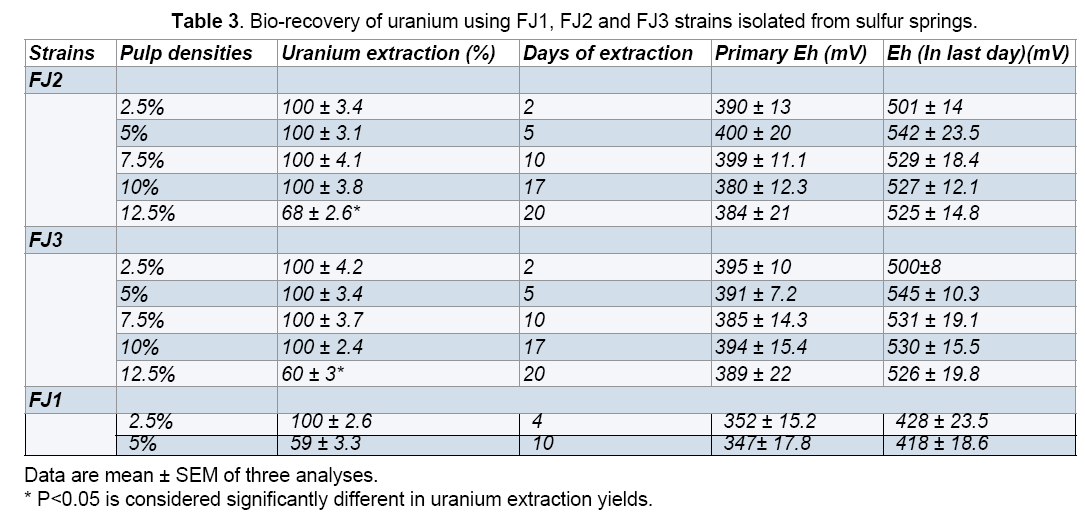
In every experiment,the variations of Eh value under different pulp densities indicate that Eh value is generally increasing,which was triggered by oxidation potential,either by a chemical or microbiologically mediated mechanisms. In addition,the results indicated that,oxidation potential varied for bioleaching pulp densities had an influence on bacterial growth as well as metal dissolution rate (Table 3).
Analyses of the leach liquor compositions
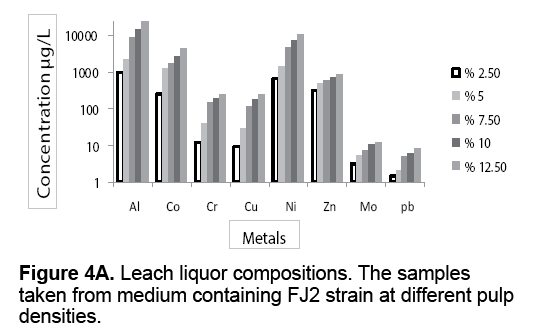
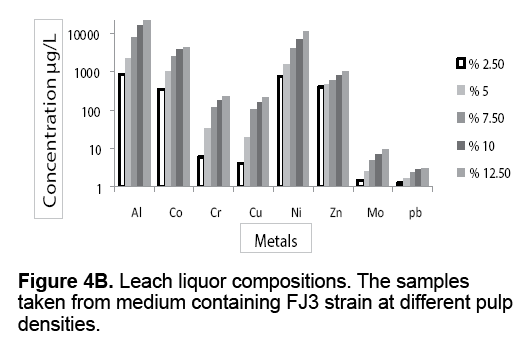
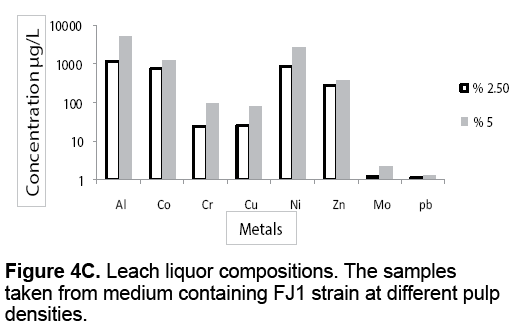
Figure 4 A-4C presents the leach liquor compositions taken from medium containing different pulp densities. The leach liquors were contained valuable metals such as Al,Co,Cr,Cu,Ni,Zn,Mo and Pb. From results,it is evident that there is high proportion of Al and Ni compared to other analyzed metals. Co is the next element followed by Zn,Cr and Cu in that order. As shown in Figure 4 A-4C,the metal leaching concentration increased with increasing pulp densities. The leaching concentration of Al and Ni were 993 μg/L and 648 μg/L at 2.5% (w/v) pulp density rising up to 23050 μg/L and 10290 μg/L at 12.5% (w/v) of pulp density,in the presence of FJ2 strain.
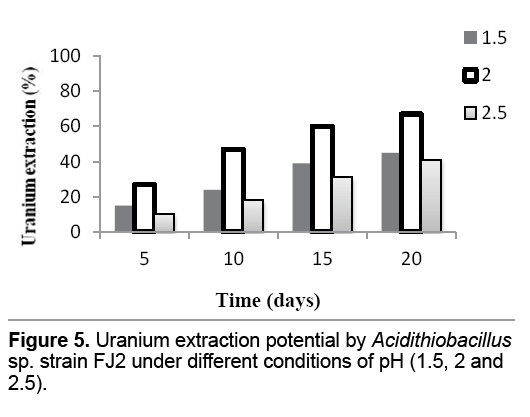
Optimization of initial ferrous iron concentration and pH
The results in Figure 5 showed that the maximum uranium extraction (67%) was achieved at pH 2,using most efficient native bacterium (Acidithiobacillus sp. strain FJ2). Acidity beyond 2 has an adverse effect on the leaching efficiency of the isolate indicating lower uranium yields (41%). It is also found that decrease the pH to 1.5 will decline the efficiency of bacterium in support of the uranium extraction yields to 45% (Figure 5 ).
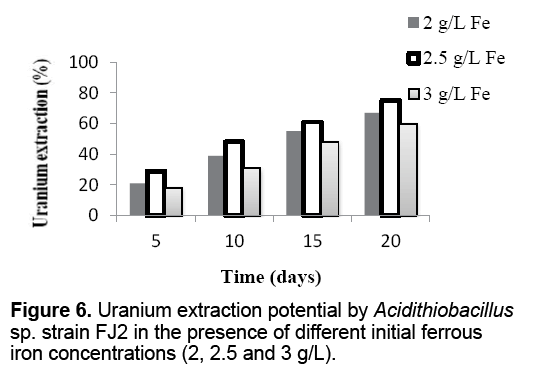
In addition,the effect of a various iron concentrations on the activity of native bacteria has been reported in Figure 6 . It has been observed that there is the highest uranium extraction yield (75%) at 2.5 g/L Fe2+ concentration. Whereas,lower uranium extraction (67 and 60%) was observed at 2 and 3 g/L of Fe2+,respectively.
Discussion
In the present study,we isolated and characterized native Acidithiobacillus sp. strains from sulfur springs Ramsar,Iran. In following,we evaluated the effectiveness of the native isolated bacteria in bioleaching optimized with different initial concentrations of ferrous iron and pH.
Our phenotypic characteristics of the isolated native bacteria indicated motive,Gram negative and rod shape bacteria (Table 2) similar with other researches [16,17]. In addition,isolated strains are member of genus Acidithiobacillus. It should be noted that since their natural habitats are ecologically extremely diverse,different Acidithiobacillus strains of the same species developing in various ecological niches,are characterized by differences in growth rate,tolerance to heavy metal ions and activities of ferrous iron and/ or sulfide mineral oxidation [18,19]. Jonhson et al in 2002 suggested that the bacteria in the same group are different strains of a single species rather than different species [20]. So,the signature sites in 16S rRNA sequences can be used as a molecular marker to distinguish the subspecies [21]. Our genotypicale consideration indicated that the isolated bacteria in the same group have 99% similarity with Acidithiobacillus sp. SM-2,Acidithiobacillus ferrooxidans ATCC 23270 and Acidithiobacillus ferrooxidans ATCC 53993.
As shown in other studies [22,23],during the incubation periods of bacteria,increase in Eh and decrease in pH values were observed in mediums (Figure 1A and 1B). Eh rising during the incubation period (Figure 1B) were triggered by the oxidation of ferrous iron/or sulfur and a chemical or microbiologically mediated mechanism [22]. As well,in 9k mediums,an initial increase in pH,attributed to the chemical dissolution of the concentrate observed between 1 and 2 days following medium pH which was declining after 2 days incubation. Several reasons may be accounted for this: generation of ferric ions from ferrous ions by bacteria,hydrolysis of ferric ions (reaction 1),sulfuric acid production,a precipitation of jarosites (reaction 2) and the chelating action of EPS which contains –OH and –COOH to release H+ into the solution (reaction 3) [23-25]:
Fe3+ + H2O → Fe (OH)3 + H+ (1)
3Fe3+ + X+ + 2HSO4- + 6H2O→ XFe3(SO4)2(OH)6 + 8H+ (2)
Where X= K+,Na+,NH4+ and H3O+
Fe3+ + 3EPS − H →Fe (EPS)3+ 3H+ (3)
Recent studies reported that the bacteria of the genus Acidithiobacillus have been widely studied for bioleaching,biodegradation and desulfurization of heavy metals [9,26-28]. So,in this study,the bioextraction ability of these bacteria was considered at different pulp densities (2.5 to 12.5% (w/v)) of low grade uranium ore sample. The results showed that,all of the isolated bacteria are capable of uranium extraction. Whereas,the bioleaching yields was diminished concomitant with increasing of pulp densities (Table 3). This may have occurred due to enhancing of inadequate diffusion of oxygen which may lead to the decelerated bacterial growth or improper growth of bacteria in higher pulp density [29]. Additionally,in bioleaching system,the accumulated metal ions became toxic for bacteria beyond certain concentrations [30]. Ginn et al. have proposed that the toxic effects of heavy metals on the growth of bacteria are not just a function of the heavy metal concentration,but they are also dependent on the cumulative contact time [31]. Similarity,in our study (Figure 4 A-4C) analysis of leach liquor indicated that,concentration of metals in leach liquors raised with increasing pulp densities. As a stress with potentially multiple impacts on different cellular components,the effects of high concentrations of metals are complex and discerning the role of a few physiological parameters [32]. The pronounced strain phenotypic and genetic heterogeneity described in A. ferrooxidans is also expressed in terms of different levels of metal resistance [33]. These differences have been attributed to several cellular characteristics such as plasmid content,and chromosomally mediated resistance mechanisms. However,the initial selection of strains that are naturally more tolerant to metals like FJ2 strain considered here,or other native strains reported by Mykytczuk et al. and Das et al.,could more improve the development of metal tolerance in A. ferrooxidans [32,34].
Furthermore,our study indicated that iron and sulfur oxidizing bacteria is more efficient than sulfur oxidizing bacteria in uranium bioleaching process (Table 3). Other studies confirmed this result indicating different effectiveness of these species in their ability for metals extraction [35,36]. The effective bioleaching of metal from ores depends upon the bacterial oxidation of Fe2+ and sulfur into Fe3+ and H2SO4,respectively [35].
It is clear that improvement in the tolerance of these bacteria to heavy metals and other inhibitory substances is of particular practical importance. As,A. ferrooxidans is a chemolithotrophic and acidophilic bacterium used inorganic compounds (iron or sulfur) as an energy source,ferrous iron concentration and pH has profound effect on the activity of bacteria in bioleaching process. As a result,in present study we optimized pH and initial concentration of ferrous iron on bioleaching process.
The results indicated that the uranium extraction rate was low at pH 1.5 and 2.5 whereas it peaked up at pH 2.0 (Figure 5 ). This defined that the acidity of the culture medium shows a negative effect on the oxidation of ferrous iron and consequently uranium extraction as biofilm formation is insignificant at pH 1.5 and 2.5 [36]. In addition,at pH above 2,the formation of jarosite hinders the iron oxidation to some extent. Jarosite creates a physical barrier between ferrous and the bacteria surface,which hinders the electron transfer from iron as well as proton diffusion from bacterial cells [37]. Moreover,decrease in the uranium extraction yields was observed when the ferrous iron concentrations were 2 and 3 g/L. While,initial ferrous iron concentration of 2.5 g/L led to the maximum uranium extraction yields (Figure 6 ). Studies indicated that with the dissolution of minerals,the excess of extracted iron ions has a negative effect on the uranium dissolution rate. During bioleaching process,Jarosite formed gradually,and coated on the surface of the remaining unreacted ore. This jarosite precipitation restricted the leaching of uranium [38].
Conclusion
In this study,we isolated the native acidophilic iron and sulfur oxidizing bacteria from sulfur springs in Ramsar province,Iran. According to the physiological features,biochemical properties and gene sequence analysis,the isolates falls into the Acidithiobacillus sp. Our results indicate that the native isolates show ability for uranium bioleaching process through 100% uranium extraction efficiency in 2.5,5,7.5 and 10% pulp densities regarding Acidithiobacillus sp. strains FJ2and 3. Whereas,Acidithiobacillus sp. strain FJ1 leaded to 100% uranium extraction at 2.5% pulp density only. In addition,the maximum uranium extraction yields were found at 2.5 g/L of initial ferrous iron concentration and an initial pH of 2.
References
- Willner J,Fornalczyk A. (2013). Extraction of metals from electronic waste by bacterial leaching. Environment Protection Engineering. 39: 197-208.
- Kalinowski BE,Oskarsson A,Albinsson Y,et al. (2004). Microbial leaching of uranium and other trace elements from shale mine tailings at Rastand. Geoderma. 122: 177-194.
- Choi MS,Cho KS,Kim DS,Ryu HW. (2005). Bioleaching of uranium from low-grade black schists by Acidithiobacillusferrooxidans. World J Mic Biotech21: 377–380.
- Bosecker K. (1997). Bioleaching: metal solubilization by microorganisms. FEMS microbial Rev. 20: 591-604.
- Abhilash S,Mehta KD,Kumar V,Pandey DB,Tamrakar KP. (2011). Bioleaching – An alternate uranium ore processing technology for India. Energy Procedia. 7: 158–162.
- Ownby DR,Newman MC. (2003). Advances in quantitative ion character-activity relationships (QICARs): using metal-ligand binding characteristics to predict metal toxicity. QSAR & Combinatorial Science. 22: 241-246.
- Johnson D,Hallberg K. (2003). The microbiology of acidic mine waters. Research in Microbiology. 154: 466-473.
- Grigorii K,Turova TP,Kondrat’eva TF,et al. (2003). Phylogenetic heterogeneity of the species Acidithiobacillusferrooxidans. International Journal of Systematic and Evolutionary Microbiology53: 113–119.
- Rawlings DE,Johnson DB. (2007). The microbiology of biomining: development and optimization of mineral-oxidizing microbial consortia. Microbiology. 153: 315–324.
- Gomez C,Bl-Zquez ML,Ballester A. (1999). Bioleaching of a spanish complex sulphide ore bulk concentrate. Minerals Engineering. 12: 93-106.
- Starkey RL,Collins VG. (1923). Autotrophs. In: Methods in Microbiology,J. R. Norris,D. W. Ribbons (Eds),New York: Academic Press. 38: 55-73.
- Silverman MP,Lundgren DG. (1959). Studies on the chemoautotrophic iron bacterium FerrobacillusferrooxidansI. An improved medium and a harvesting procedure for securing high cell yields. J Bacteriol. 77: 642–647.
- Buchanan RE,Gibbons NE. (1974). Bergey’s of determinative bacteriology,America: United States of America. 529-563.
- Edwards U,Rogall T,Blocker H,Emde M,Bottger EC. (1989). Isolation and direct complete nucleotide determination of entire genes. Nucleic Acids Res. 17: 7843–7853.
- Golmohammadi H,Rashidi A,Safdari SJ. (2012).Simple and rapid spectrophotometric method for determination of uranium (VI) in low grade uranium ores using arsenazo(III). Chemistry and Chemical Technology. 6: 245-249.
- Ryu HW,Moon HS,Lee EY,Cho KS,Choi H. (2003). Leaching Characteristics of Heavy Metals from Sewage Sludge by Acidithiobacillusthiooxidans.MET,J. Environ. Qual. 32: 751-759.
- Johnson DB,Okibe N,Roberto FF. (2003). Novel thermoacidophilic bacteria isolated from geothermal sites in Yellowstone National Park: physiological and phylogenetic characteristics. Arch. Microbiol. 180: 60– 68.
- Ageeva SN,Kondrat’eva TF,Karavaiko GI. (2001). Phenotypic characteristics of Thiobacillusferrooxidans strains. Mikrobiologiia,70: 226–234.
- Kondratyeva TF,Pivovarova TA,Muntyan LN,Karavaiko GI. (1999). Strain diversity of Thiobacillusferrooxidans and its significance in biohydrometallurgy,In Biohydrometallurgy and the Environment toward the Mining of the 21st Century. 89–98.
- Johnson DB,Bridge TA. (2002). Reduction of ferric iron by acidophilic heterotrophic bacteria: Evidence for constitutive and inducible enzyme systems in Acidiphiliumspp. Journal of Applied Microbiology. 92: 315-321.
- Bertil P,Goran B,Francois T,Mathias U.,Karlerik J. (1998). Molecular evolution of mycoplasma capricolum subsp. capripneumoniaestrains,based on polymorphisms in the 16S rRNA genes. J. Bateriol,180: 2350-2358.
- MeruaneG,Vargas T. (2003). Bacterial oxidation of ferrous iron by Acidithiobacillusferrooxidans in the pH range 2.5–7.0. Hydrometallurgy. 71: 149–158.
- Liang G,Tang J,Liu W,Zhou Q. (2013). Optimizing mixed culture of two acidophiles to improve copper recovery fromprinted circuit boards (PCBs). J HazardMater,15: 250–251.
- Gomez E,Ballester A,Gonzalez F,Blazquez ML. (1999). Leaching capacity of a new extremely thermophilic microorganism,Sulfolobusrivotincti. Hydrometallurgy. 52: 349–366.
- Yu RL,Tan JX,Gu GH,Hu YH,Qiu GZ. (2010). Mechanism of bioleaching chalcopyrite by Acidithiobacillusferrooxidansin agar-simulated extracellular polymeric substances media. J Central South University Techno.17: 56-61.
- Yang Y,Chen S,Li S,Chen M,Chen H,Liu B. (2014). Bioleaching waste printed circuit boards by Acidithiobacillusferrooxidans and its kinetics aspect. J Biotechnol.10:24-30.
- Wang J,Zhu S,Zhang Y,et al. (2014). Bioleaching of low-grade copper sulfide ores by Acidithiobacillusferrooxidans and Acidithiobacillusthiooxidans. Journal of Central South University. 21: 728-734.
- Peng H,Yang Y,Li X,et al. (2006). Structure analysis of 16S rDNA sequences from strains of Acidithiobacillusferrooxidans. J. Biochem. Mol. Biol. 39: 178–182.
- Debabrata P,Dong J,Kim J,Seoung W. (2010). Microbial Leaching Process to Recover Valuable Metals from Spent Petroleum Catalyst Using Iron Oxidizing Bacteria. World Academy of Science,Engineering and Technology. 4: 02-22.
- Mei Li H,Jun Ke J. (2001). Technical note in influence of Cu+2 and Mg+2 on the growth and activity of Ni+2 adapted thiobacillusferrooxidans. Miner Eng.14: 113-116.
- Ginn T,Sengor SS,Barua S,Moberly J,Peyton B. (2006). Metal toxicity effects on microbial growth and degradation,Slovenia and U.S,Workshop on Environmental Science and Engineering,Ljubljana,Slovenia. 39-40.
- Mykytczuk NCS,Trevors JT,Ferroni GD,et al. (2011). Proteomic comparisons of cold adaptation in psychrotrophic and mesophilic strains of Acidithiobacillusferrooxidans. Antonie van Leeuwenhoek.100: 259-277.
- Dopson M,Baker-Austin C,Koppineedi PR,Bond PL. (2003). Growth in sulfidic mineral environments: metal resistance mechanisms in acidophilic micro-organisms. Microbiology149: 1959-70.
- Das A,Modak JM,Natarajan KA. (1997). Studies on multi-metal ion tolerance of Thiobacillusferrooxidans. Miner Eng. 10: 743-9.
- Haragobinda S,Ashish P,Dong JK,Seoung-Won L. (2013). Comparison of Bioleaching of Metals from Spent Petroleum Catalyst Using Acidithiobacillusferrooxidansand Acidithiobacillusthiooxidans. International Journal of Chemical,Nuclear,Metallurgical and Materials Engineering. 7: 499-503.
- KoMS,Park HS,Kim KW,Lee JU.(2013). The role of Acidithiobacillusferrooxidans and Acidithiobacillusthiooxidans in arsenic bioleaching from soil. Environ Geochem Health.35: 727-33.
- Shafia F,Wilkinson RF. (1969) Growth of Ferrobacillusferrooxidans on organic matter.J. Bacteriology. 97: 256-260.
- Kaibin F,Hai L,Deqiang L,Wufei J,Ping Z. (2014). Comparsion of bioleaching of copper sulphides by Acidithiobacillusferrooxidans. Environ Geochem Health. 13: 664-672

Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences