Electrophoretic Characterization and the Relationship between Some Brassica Species
Munazza Sadia, Salman A. Malik, Malik Ashiq Rabbani, S.R. Pearce
1Department of Biochemistry,Quaid-I-Azam University,Islamabad,Pakistan
2Institute of Agri-Biotechnology & Genetic Resources,NARC,Islamabad,Pakistan
3School of life sciences,University of Sussex,Brighton,UK
Abstract
The objective of this study was to assess the genetic divergence available in different genotypes of oilseed Brassica based on analyses of seed storage proteins, for the identification of genetically diverse and agronomically superior genotypes of Brassica seed, which may generate putative transgressive segregates on hybridization. Diversity within species of Brassica can be analyzed at molecular level with the help of seed storage protein by using SDS-PAGE. On the basis of banding pattern zymogram (diagrammatical representation of different protein bands) were sketched, by which molecular weight of specie specific bands were calculated on the basis of Rf values of the bands on the gel. Through statistical analyses dendrogram was formed and genotypes were clustered into different groups by applying UPGMA (unweighed pair group mean analyses).
https://matadorbet-giris.com https://grandpashabeti.com https://betiste.com https://bahsegelgirisi.com https://klasbahisgirisi.com https://jasminbeti.com https://hepsibahise.com https://dinamobetgirisi.com https://betvolegirisi.com https://betpark-girisi.com https://betlikegiris.com https://betboogirisi.com https://sultanbetegiris.com https://tulipbetgir.com https://padisahbetegir.com https://savoybettinge.com https://goldenbahisegiris.com https://maksibetgirise.com https://fenomengiris.com https://jojobetgiris.xyz https://tarafgiris.com
Keywords
Brassica,diversity,SDS-PAGE,seed protein.
1. Introduction
Brassica species are used as oil seed crops (B.napus and B.juncea),leafy vegetables and turnip (B.oleracea and B.rapa),and are cultivated worldwide. Especially,in East Asia,many varieties of B.rapa are used as agronomically important vegetables. In general,genetic improvement of crops can be accelerated when broad genetic diversity and the information of these genetic resources are available. Research on Brassica germplasm could enhance the edible oil production and nutritional benefits of these crops. The collection of these genetic resources and the assessment of genetic diversity within and between landraces should have priority for varietal improvement. At the same time it is necessary to develop better methods of characterization and evaluation of germplasm collections,to improve strategies for conservation and collection of germplasm and to increase the utilization of plant genetic resources. The electrophoresis of seed storage protein is a method to investigate genetic variation and to classify plant varieties [1]. Seed protein is not sensitive to environmental fluctuations; its banding pattern is very stable which advocated for cultivars identification purpose in crop. It has been widely suggested that such banding patterns could be important supplemental method for cultivars identification,particularly when there are legal disputes over the identity of a cultivar or when cultivars are to be patented [2]. Seed storage protein is useful tool for studying genetic diversity of wild and cultivated rice [3]. However,the information on the SDS-PAGE on different species of Brassica for genetic diversity is still limited [4]. Analyses of SDS-PAGE are simple and inexpensive,which are added advantages for use in practical plant breeding. In this study,a survey of seed protein was carried out to (a) assess the protein polymorphisms within and different cultivated species of Brassica (b) clarify the genetic nature of polymorphic bands.
2. Methods
Plant materials
Altogether,30 cultivars of Brassica rapa,Brassica juncea,Brassica carinata and Brassica napus were used for this experiment (Table 1).

Electrophoresis
A single seed was grounded with a mortar and pestle and 10mg (0.01g) out of this seed flour was taken into a 1.5ml micro-tube. 400μl of the protein extraction buffer (62.5mM Tris-Hcl,pH 6.8,2% SDS,10% glycerol,5% b-mercaptoethanol,5 M urea and 0.0001% bromo-phenol blue) was added and mixed well by vortexing. The crude homogenates were then centrifuged at room temperature with 15000rpm for 10 min. Thereafter,6.5μl of the extract was directly analyzed by SDS-Polyacrylamide gel electrophoresis (SDS-PAGE) using 15% (w/v) mini slab gel. Electrophoresis was carried out at 20V for stacking gel and 80V for separation gel in a buffer solution containing 0.025M Tris,0.129M glycine and 0.125% SDS,until the dye front head migrated to within 2mm of the end of the gel. Gels were stained with 0.5% coomassie brilliant blue (CBB) G-250 in acetic acid-methanol-water (3:22:25 volume ratio) for two hours and destained in acetic acidmethanol- water (5:20:75 volume ratio) for over night. Banding patterns were scored from at least two electrophoregrams for each cultivar.
Evaluation
Gels were evaluated by eye on a light box. Determination of the apparent molecular weight of individual protein subunits was carried out using molecular weight marker proteins: b-galactosidase,116.0 KD; Bovine serum albumin,45.0 KD; Lactate dehydrogenase,35.0 KD; REase Bsp 981,14.4 KD. (Protein molecular wt. Marker,Fermentas),applied on each gel and Rf values,the relative mobility of protein subunits was calculated.
Statistical analyses
All the monomorphic and polymorphic bands visible to the eye were scored and only unambiguously scored bands were used in the analyses. Each band was given score of 1 for presence or polymorphism and 0 for absence. Data analyses were conducted using NTSys-pc,version 2.2 (Exeter soft ware,Setauket,N.Y.). Similarities between cultivars were estimated using Dice coefficient of similarity (Rohlf 1992). Cluster analyses were conducted on similarity estimates using the unweighed pair- group method for arithmetic averages (UPGMA) and the resulting clusters were expressed as dendrograms.
3. Results and Discussion
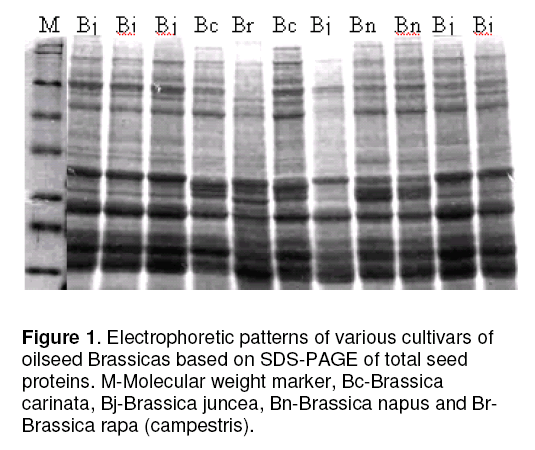
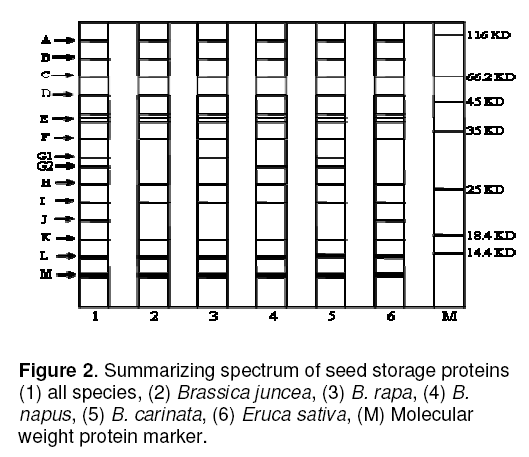
The protein patterns of 30 Brassica cultivars after SDS-PAGE are shown in Figure 1. In total 29- 31 bands per cultivars were detected in electrophoregrams. Of these polymorphic bands appear at 13 positions designated as A,B,C,D,E,F,G,H,I,J,K,L and M respectively (Figure 2 ). Bands at position ‘E’ were divided into three patterns (E1,E2 and E3). These showed polymorphism on the basis of difference in protein intensity among genotypes. Bands at positions G and J showed presence- or- absence type polymorphism. At position G,some genotypes expressed a single band and others showed a pair of bands and two levels of mobility of the bands were detected. These banding patterns were recognized in the position as G1 and G2 respectively. B. rapa exhibited a specie-specific band at position G1 (M.wt-32.6 KD),B. napus produced a specie-specific band at G2 (M.wt- 30.7KD),B. carinata exhibited both of these bands at positions G1 and G2 where as B. juncea possessed none of these. Eruca sativa species exhibited a characteristic band at position ‘J’ (M.wt- 20 KD).
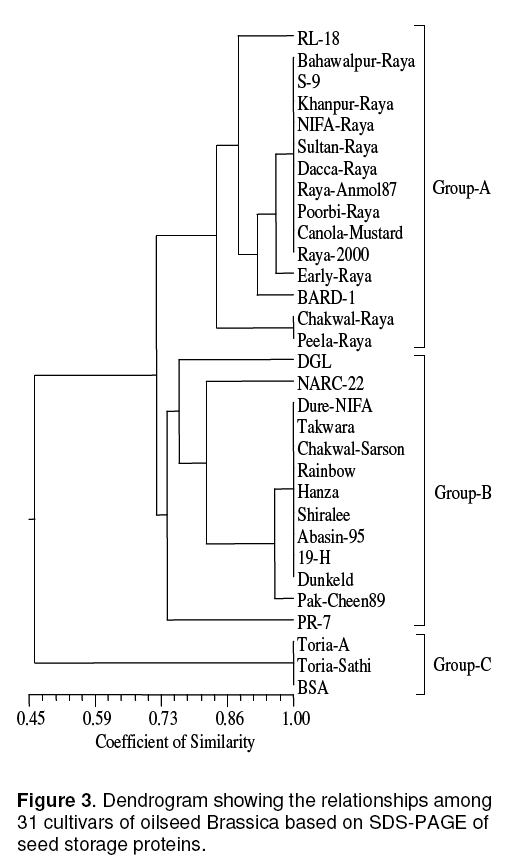
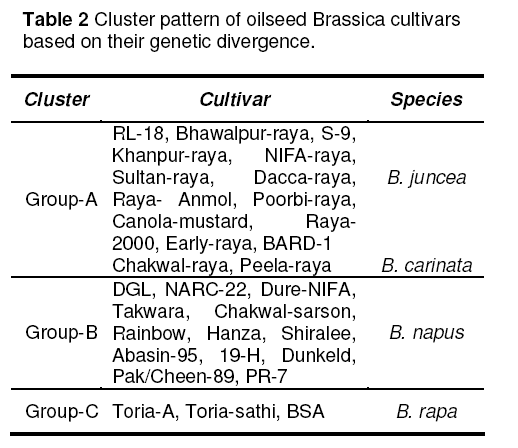
Three groups or clusters obtained from Dendrogram,clustering of population follow their genetic similarity (Figure 3 ). Cluster 1 comprises of fifteen genotypes where as cluster 2 comprises thirteen and cluster 3 comprises of three genotypes (Table 2). In this study intra and inter specific variation of seed storage proteins of 30 cultivars of oilseed Brassicas from Pakistan were analyzed. No report on SDSPAGE of seed protein of Brassica varieties of Pakistan is available,so it seems to say that SDSPAGE technique has proven to be a useful tool in supporting classical taxonomy studies [5]. It was possible to distinguish certain genotypes based on seed protein. Landraces are a useful source of genetic variation and the greater the variation,the greater the chances of a landrace possessing genes of gene combinations of interest to plant breeders [6]. The high stability of seed protein profile and its additive nature make seed protein electrophoresis a powerful tool in elucidating the origin and the evolution of cultivated plants [7].
4 Conclusions
The present investigation revealed high variation in different cultivars of oil seed mustard in Pakistan with regard to their total seed protein profiles. Regarding interspecific variation among cultivars this investigation revealed no variation. The genetic affinities within cultivars of same species generally corroborated the morphological analysis [8]. Phenotypically,most of the oilseed cultivars belonging to same specie also showed close association. This uniformity in interspecies cultivars also agreed with the findings of Ladizisky and Alder [9] and Ahmad and Slinkard [10],who examined different cultivars of cultivated chickpea and concluded that seed protein was a very conservative trait in chickpea. Similarly,Raymond et al. [11] and de vries [12] also reported similar electrophoretic patterns of protein among the cultivars of sunflower and lettuce,respectively.The result of differentiation of yellow sarson and brown seeded types of Brassica shows a similar agreement with the report of Das et al. [13] where the result clearly separated the yellow seeded and brown seeded varieties.
Acknowledgements
We acknowledge with thanks to Dr. Zahoor Ahmed and Dr. Shahid Masood for extending lab facilities of Plant Genome Research Institute,NARC,Islamabad.
References
- Isemura T.,Shiyo N.,Shigeyuki M.,et al. (2001) Genetic variation and geographical distribution of Azuki bean (Vigna angularis) landraces based on the electrophoregram of seed storage proteins. Breeding Sci.,51: 225-230.
- Tanksley S.D.,Jones R.A. (1981) Application of alcohol dehydrogenase allozymes in testing the genetic purity of F1 hybrids of tomato. Hort. Sci.,16: 179-181.
- Thanh V.O.C.,Hirata Y. (2002) Seed storage protein diversity of three rice species in the Mekong Delta. Biosphere Conservation,4: 59-67.
- Rahman M.M.,Hirata Y. (2004) Genetic diversity in Brassica species using SDS-PAGE analysis. J. Biol. Sci.,4(2): 234-238.
- Thanh V.O.C.,Nguyen T.N.,Hirata Y.,et al. (2003) Antenna protein diversity of prawns (Macrobrachium) in the Mekong Delta. Biosphere Conservation,5: 11- 17.
- Damania A.B.,Porceddu E.,Jackson M.T. (1983) A rapid method for the evaluation of variation in germplasm collections of cereals using polyacrylamide gel electrophoresis. Euphytica 32: 877-883.
- Ladizinsky G.,Hymowitz T. (1979) Seed protein electrophoresis in taxonomic and evolutionary studies. Theor. Appl. Genet. 54: 145-151.
- Rabbani M.A.,Iwabuchi A.,Murakami Y.,et al. (1997) Phenotypic variation and the relationships among mustard (Brassica juncea L.) germplasm from Pakistan. Euphytica,101:357-366.
- Ladizinsky G.,Adler A. (1975) The origin of chickpea as indicated by seed protein electrophoresis. Isr. J. Bot. 24: 183-189.
- Ahmad F.,Slinkard A.E. (1992) Genetic relationships in the genus Ciser L. as revealed by polyacrylamide gel electrophoresis of seed storage proteins. Theor. Appl. Genet. 84: 688-692.
- Raymond J.,Inquello V.,Azanza J.L. (1991) The seed proteins of sunflower: comparative studies of cultivars. Phytochemistry 30: 2849-2856.
- De Vries I.M. (1996) Characterization and identification of Lactuca sativa cultivars and wild relatives with SDS-electrophoresis (Lactuca sect. Lactuca,Compositae),Genetic Resources and Crop Evolution 43: 193-202.
- Das S.,Mukherjee K.K. (1995) Comparative study on seed proteins of Ipomoea. Seed Sei. & Technol. 23: 501-509.

Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences