A Study of Comparing Changes of Glu-Alx Promoter Sequences in Iranian Diploid and Hexaploid Wheat
Nastaran Partovi, Mohsen Ebrahimi, Ali Izadi darbandi, Hosseinali Ramshini
Nastaran Partovi*, Mohsen Ebrahimi, Ali Izadi darbandi, Hosseinali Ramshini
Department of Agronomy and Plant Breeding sciences, College of Aburaihan, University of Tehran, Tehran, Iran.
- Corresponding Author:
- E-mail: mebrahimi@ut.ac.ir
Received Date: Februaury 27, 2016; Accepted Date: May 27, 2016; Published Date: June 03, 2016
Citation: Partovi N, Ebrahimi M, Darbandi AI, et al. A Study of Comparing Changes of Glu-Alx Promoter Sequences in Iranian Diploid and Hexaploid Wheat. Electronic J Biol, 12:3
Abstract
The rise of world population and food crisis in developing countries have directed the attention of agriculture experts to the most strategic crops, namely, wheat in the world. Low quality of wheat bread is the most important factor in its wastes, thus it seems to be essential to invest on functional studies in this field. Genetic changes for the purpose of improving the baking quality of wheat bread and its endosperm construction require desirable gene promoters so as to increase tragenes expression, so separating glutenin gene promoters with high molecular weight (Glu-Alx) from diploid and hexaploid wheat was examined in this study. Designing proper primers and applying them on the genomic DNA were conducted in polymerase chain reaction (PCR) and promoter fragments as long as 1100 bp were produced. Then, the fragments were transmitted to the pTG19 vector and white colonies were evaluated through enzyme digestion. Results showed that the desired fragment was found in the studied colonies. Online BLAST analysis and ClustalW multiple alignment indicated that there were many sequences similarity of this sequence with other known HMWGS promoter sequences. Generally the results showed that the homologized promoter can be used in molecular euphonic programs of wheat quality and it can be used in making expression constructions and gene transfer or its overexpression.
Keywords
Promoter; High-molecular-weight glutenin; Diploid and Hexaploid Wheat
1. Introduction
Wheat is among the most adaptable corns and accounts for about 90% of the cultivated areas in the world [1]. Unfortunately this strategic corn are wasted in different ways which low quality of produced flours is one of the reasons for bread wastes. The quality of wheat glutenin depends on genotypes of wheat varieties and this is the trait making wheat flours prominent. The amount of wheat baking quality is controlled with high molecular weight glutenin (HMW-GS) and low molecular weight glutenin (LMWGS) do not have a significant effect on the quality [2]. Genes controlling HMW-GS are encoded by gene loci of Glu-1 in three sites including Glu-A1, Glu-A2, and Glu-D1 which are located on long arms of 1A, 1B, and 1D chromosomes and each of these gene loci comprises two closely related genes named as Glu1-1 and Glu1-2 coding x-type and y-type subunits in terms of molecular weight [3]. The genetic transformation provides a means of introducing the novel genes for modifying the cereal grain characteristics. For example, the transgenes can be used to alter the relative amounts of the specific proteins or starch components to make new or improved processed products. To this aim, gene promoters of wheat HMW-GS which support high level of prolamins synthesis through the linear phase of seeds maturity are regarded as the best choice for expressing transgenes in the endosperms of corns. These promoters are among organ and tissue specific promoters and cause expression of the genes in seeds. Promoters areas of HMW-GS genes and wheat gliadins, barley hordeins and sekalins have a section called endosperm box containing two neighboring sections, namely, GCN4-motif section (G section or N section) and endosperm or prolamine section (E section). The two-section endosperm box is located at the distance of 300 bp above the encoding area. The section E is required to express prolamine genes in corn, barley and wheat endosperms and tobacco cotyledons. GCN4-motif section in ferments and other organisms have been created in response to the environment and feeding conditions and it is guessed that sections similar to this section are also found in wheat prolamine genes, barley C-hordein and alpha Zein in corn (in response to nitrogen) [4]. In Furtado et al. [5] the gene promoter of high molecular weight glutenin subunits (HMW-GS) with 425 bp in length in transgenic wheat caused gene reporter to be expressed in endosperm and aleurone layer and not to be expressed in leaves and roots. Lamacchia et al. [4] studied endosperm specific promoter in transgenic durum wheat showing that expression was higher in the central starchy endosperm cells with no expression in the aleurone cells. Ayyari et al. [6] cloned low molecular glutenin subunits specific promoter in durum wheat (Triticum durum). Shahmoradi et al. [7] performed cloning promoter specific low molecular glutenin subunits of Iranian wheat. The study of the sequences cloned with BLAST and clustalW software programs showed that these sequences had many similarities to other well-known storage proteins. Weibo used Nested PCR method and specific initiators to study molecules and clone HMW promoter and amplified a 1050 bp promoter fragment. The shortage of gene promoters suitable for transgenes expression in corns endosperm is regarded as a massive restriction on achieving the required mount and pattern of gene expression. So it is very important to study and identify Glu-Alx gene sequences in these plants. Therefore, cloning Glu- Alx gene and also studying differences among the promoters in the two Iranian diploid and hexaploid varieties of wheat were investigated in this study.
2. Materials and Methods
2.1 Plant materials a DNA extraction
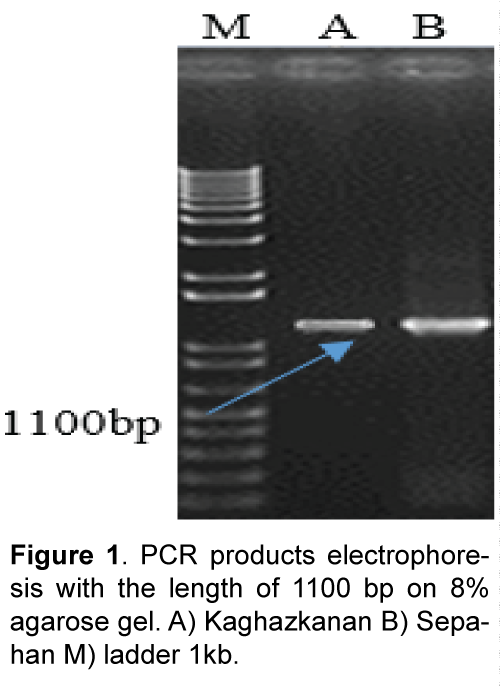
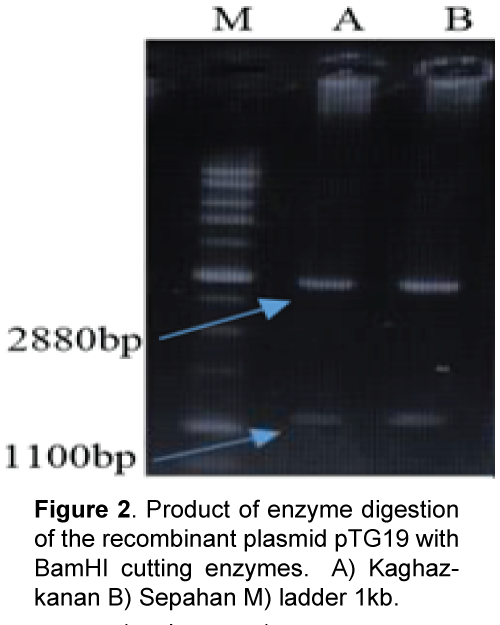
Plant materials used in this study consist of diploid wheat (T. boeoticum) Kaghazkanan varietie and hexaploid wheat (T.aestivum) Sepahan varietie which were supplied from gene bank of the Institute for Breeding and Producing Seeds and Seedlings. The seeds were planted in vases and then young leaves were used. DNA was extracted using a modified Saghai et al. [8] method. DNA quality and quantity were measured using spectrophotometry and agarose gel electrophoresis. Direct and reverse primers were designed with the software Primer3 referring to nucleotide bank information (NCBI) and two sweep primers used for each varietie in PCR reaction (Table 1). Amplification of DNA fragments in polymerase chain reaction with a final volume of 25 μl (PCR) was performed containing 2.5 μl PCR buffer with 10 x concentration, 0.5 μl dNTP with 10 μM, 2 μl magnesium chloride with a concentration of 50 μM, 2 μl of each primer with 5 μM concentration, and 0.5 μl taq polymerase enzyme with a concentration of 5 units per μl were reached to the concentration of 15 μl with double distilled water and then adding 2.5 μl DNA with a concentration of 25 ng/μl it was reached to the final volume Polymerase chain reaction steps included an initial denaturation at 94°C for 5 min, 35 cycles of denaturation at 94°C for 40 seconds, connecting at 60°C for 1 min and synthesis and extension at 72°C for 1.5 min and final extension at 72°C for 7 min. In order to study the PCR product, 8% agarose gel electrophoresis, stained with ethidium bromide and photographed under UV light was used (Figure 1). Isolation of the target fragment from agarose gel was performed using Silica Bead DNA Gel Ext kit. Cloning and sequencing DNAs: PCR product was attached to the pTG19 vector with T4-ligase enzyme. After preparing E. coli – competent cells of DH5a variant at OD600 = 0.4 based on alkali calcium chloride method [9], the recombinant plasmid was transferred using a thermal shock to the bacteria [10] Transgenic bacterial cells were selected using blue-white screening and after cultivating transgenic bacteria, recombinant plasmids were extracted based on alkaline method with SDS. One microgram (μg) of recombinant plasmids was digested using BamHI restriction enzyme and the reaction product was studied applying 1.5% agarose gel electrophoresis (Figure 2). The accuracy of sequences cloned through DNA sequencing was conducted by Bioneer Company using M13 primers in both directions. After sequencing, the software programs BLAST and clustalW were used to study the similarities and differences of the studied plant materials. Predicting position of the promoters and transcription initiation codon were performed with Neural Network Promoter Predictions software. Analysis for cic-acting response elements was done with PlantCARE [11] and PLACE [12] databases.
| Primer sequences | Varietie |
|---|---|
| AGGGAAAGACAATGGACA | Kaghazkanan |
| GTCTCGGAGTTGCTGGTC | |
| ACACTTCATCATCTTCCTCG | Sepahan |
| CAGATTTAATTGTCCATTA |
Table 1. List of used primers.
3. Results and Discussion
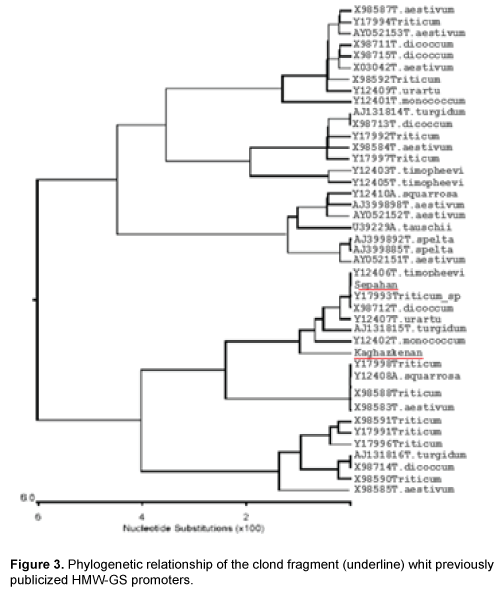
Results of polymerize chain reaction, using HMWPrimer GS gene promoter in two diploid and hexaploid varieties of wheat led to the amplification of 1100 bp. Phylogenetic analysis using similar sequences in Nucleotide Information Bank (NCBI) and also the sequences obtained from cloned fragments was conducted to determine that cloned promoters fragments belong to which gene high molecular weight glutenin subunit (Figure 3). Results from analysis indicate that promoters separated from two diploid and hexaploid wheat species are related to Glu-Alx gene promoter. As the two promoters separated from bread and boeoticum wheat are related to Glu-Alx gene and are paced in the same group, so ClustalW software was used to place the sequences in the same row. Results indicated that the two mentioned promoters had wide differences with each other. Kaghazkanan species promoter sequence also was compared to other gene promoters of Glu-Alx in different varieties of wheat and the results indicated that the two sequences had 99% of similarity and 60 sequences had 89% of similarity. The studies on the gene promoter of Sepahan varietie indicated that 25 sequences had 99 percentages of similarity and 55 sequences had 89 percentages of similarity. Being placed in the same row in diploid species of Kaghazkanan wheat showed that about 137 bp at the end 3' of this fragmentation are correlated with 143 bp at the end of 5' gene and regarding Sepahan varietie about 123 bp at the end of 3' of this fragment are correlated with 127 bp of 5' gene. This issue suggests that separated sequences are placed above Glu-Alx gene.
Starting point (+1) in Kaghazkanan varietie was predicted by Neutral Network Promoter Predictions as 67 bp above translation initiation codon. This fragment comprising 960 bp is higher translation initiation codon and contains important cis-acting response elements. The most important cis-acting elements and their function are presented in Table 2. The position of indicated cis-acting elements in the Glu-Alx gene of Kaghazkanan varietie is showed in Figure 4.
| Function | cis-acting element |
|---|---|
| cis-acting element involved in the abscisic acid responsiveness | ABRE |
| common cis-acting element in promoter and enhancer regions | CAAT-box |
| cis-acting regulatory element related to meristem expression | CAT-box |
| cis-acting regulatory element related to meristem specific activation | CCGTCC-box |
| cis-acting regulatory element involved in light responsiveness | G-Box |
| gibberellin-responsive element | GARE-motif |
| cis-regulatory element involved in endosperm expression | GCN4-motif |
| light responsive element | GT1-motif |
| cis-acting element involved in heat stress responsiveness | HSE |
| part of a light responsive element | I-box |
| cis-acting regulatory element required for endosperm expression | Skn-1-motif |
| core promoter element around -30 of transcription start | TATA-box |
| cis-acting element involved in salicylic acid responsiveness | TCA-element |
| cis-acting regulatory element involved in the MeJA-responsiveness | TGACG-motif |
| cis-acting regulatory element involved in circadian control | Circadian |
| Prolamin-box |
Table 2. cis-acting elements inside Glu-Alx gene of Kaghazkanan varietie.
Studies also showed that (+1) starting point in Sepahan varietie was predicted as 85 bp above translation initiation codon. This fragment comprising 893 bp is higher translation initiation codon and contains important cis-acting response elements. The most important cis-acting elements and their function are presented in Table 3. The position of indicated cis-acting elements in the Glu-Alx gene of Sepahan varietie in Figure 5.
| Function | cis-acting element |
|---|---|
| cis-acting element conferring high transcription levels | 5UTR Py-rich stretch |
| common cis-acting element in promoter and enhancer regions | CAAT-box |
| cis-acting element involved in light responsiveness | ACE |
| light responsive element | Box I |
| cis-acting regulatory element involved in light responsiveness | G-Box |
| gibberellin-responsive element | GARE-motif |
| cis-regulatory element involved in endosperm expression | GCN4-motif |
| cis-acting element involved in low-temperature responsiveness | LTR |
| light responsive element | MNF1 |
| cis-acting element involved in heat stress responsiveness | HSE |
| cis-acting element involved in defense and stress responsiveness | TC-rich repeats |
| core promoter element around -30 of transcription start | TATA-box |
| cis-acting element involved in salicylic acid responsiveness | TCA-element |
| part of a light responsive element | TCT-motif |
| cis-acting regulatory element involved in circadian control | Circadian |
Table 3. cis-acting elements inside Glu-Alx gene of Sepahan varietie.
4. Discussion
TATA-box which exists in most eukaryotic genes is located in -30 position above translation initiation codon in the two varieties. The box is located in -26 to -31 higher than translation initiation codon in eukaryotic gene promoters [13]. CAAT-box is located in positions -835, -742, -463, -347, -275, -246, and -171 in Kaghazkanan varietie and in positions +62 and -434 in Sepahan varietie. CAATbox is a common motif in most eukaryotic gene promoters which plays an important role in regulating the beginning of translation [14]. G-box is among important identified motifs which is a motif with conserved sequences (TACGTG) in most plant genes promoters and is the first identified regulatory sequence in response to light which can attach to a family of transcription factors (GBF) with bZIP domain [15,16]. It is worth mentioning that existence of regulatory motifs including box 4, G-box, GT1- motif, SP1, and I-box in Glu-Alx gene promoter which is among responsive factors to light indicates the importance of light in increasing the expression of the gene. Identified W-box motif in the promoter of Kaghazkanan varietie has attachment place for the WRKY family of transcription factors. Other important motifs were found included the A, B and C consensus sequence in the promoter of Kaghazkanan varietie. A, B and C motifs were conserved sequences concerning the cereal storing protein expression. Both A and C were conservative elements in the homogeneous sequence of the published wheat HMW-GS 5’ upstream. Furthermore, they have middle conservation with 5’ upstream different region of several kinds α/β type gliadin gene, and were predicted that they have relation to gene expression and regulation. B element was “-300 element”, whose central sequence were TGCAAAA or TCTAAAG. It existed in the 5’ conserved region of many cereal storing protein genes, including the HMW-GS and played a significant role in the endosperm-specific expression. Based on these results, it could be hypothesized that the HMW-GS gene promoter might be an endosperm-specific promoter.
These translation factors play an important role in regulating different plant activities such as defense against pathogens. TCA is another motif which is a junction of transcription factors regulating the expression of effective genes in response to salicylic acid involved in plants. Promoters of seeds accumulated proteins are parts of organ specific promoters resulting in gene expression in certain organs and determine specific expression time patterns. Due to lack of Prolamin-box, A, B and C motifs which are important for regular expression of distinctive genes of endosperms in Sepahan varietie, Glu-Alx gene was not expressed. This issue can be regarded as one of the reasons for lower quality of this varietie as compared to Kaghazkanan varietie [17] identified three important conserved motifs including GCN4-motif, Prolamin-box, and AACA motifs in distinctive promoters of wheat, barley, corn, and rice endosperms. It must be mentioned that Guillaumie et al. [18] stated that GCN4-motif and Prolamin-box constitute endosperm box. Making use of promoters which merely act in some fibers can be effective in achieving distinctive goals. Among functions of a distinctive promoter are to use them in breeding programs and produce recombinant proteins in seed tissues. Regarding the importance of wheat as the most important cultivated plants in the world, it is of high importance to separate promoters of distinctive proteins of seeds and can be used to conduct studies and qualitatively quantitatively breed it.
5. Conclusion
In the study, the promoter fragments of 960 bp and 893 bp above the translation initiation codon were segregated based on PCR method from Kaghazkanan and Sepahan varieties, respectively. As the importance of cis-acting elements located inside promoter regions, it is necessary to find and characterize functional motifs responding special elicitors to regulate gene expression in eukaryotic organisms. Subsequent analysis with bioinformatics databases such as PlantCARE and PLACE indicates that there are several functional motifs involved in specific elicitors responding e.g., light, hormones, salicylic acid etc. It is a helpful step to understand which elicitors are affecting gene expression enhancement in wheat.
References
- Gianibelli MC, Larroque OR, MacRitchie F,et al. (2001). Biochemical, genetic and molecular characterization of wheat glutenin and its component subunits. Cereal Chemistry. 78: 635-646.
- IzadiDarbandi A, Yazdi-samadi B. (2012). Marker-assisted selection of high molecular weight glutenin alleles related to bread-making quality in Iranian common wheat (Triticumaestivum L.). Genetics.91:193-198.
- Johal J, Gianibelli MC, Rahman S, et al. (2004). Characterization of lowmolecular-weight glutenin genes in Aegilopstauschii. Theoretical and Applied Genetics. 109:1028-1040.
- Lamacchia C, Shewry PR, Di Fonzo N, et al. (2001). EndospermÃÆâÃâââ¬ÃâÃÂspecific activity of a storage protein gene promoter in transgenic wheat seed. Journal of Experimental Botany. 52:243-250.
- Furtado A, Henry RJ, Pellegrineschi A. (2009). Analysis of promoters in transgenic barley and wheat. Plant biotechnology Journal.7:240-253.
- Ayyari M, BaghbanKohnerouz B, Golizade A. (2011). Cloning of High Molecular Weight Glutenin Subunit (HMW-GS) Gene Specific Promoter from Durum Wheat (Triticum durum).The 7th National Biotechnology Congress of Iran 12-14 Sep, Tehran, Iran.
- Shahmoradi A, BaghbanKohnehrouz B, doraniuliaie E. (2012). The Cloning of LMW-GS Endosperm Specific Promoter from Iranian Wheat. 12th Iranian Genetics Congress, May 22-24, 2012, Beheshti international congress center, Tehran, Iran.
- SaghaiMaroof MA, Soliman K, Jorgensen RA, et al. (1984). Ribosomal DNA spacer-lengh polymorphisms in barley: mendelian inheritance, chromosomal location and population dynamics. Proceedings of the National Academy of Sciences. 81:8014-8018.
- Matsuda J, Okabe S, Hashimoto T, et al.(1991). Molecular cloning of hyoscyamine 6β- hydroxylase, a 2- oxoglutarate- dependent dioxygenase, from cultured root of Hyoscyamusniger. Biol chem. 266: 9460-9464.
- Cohen SN, Chang AC, Boyer HW, et al.(1973). Construction of biologically functional bacterial plasmid in vitro. ProcNatlAcadSciUSA. 70: 3240-3244.
- Lescot M, Dehais P, Thijs G, et al. (2002). PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30:325-327.
- Higo K, Ugawa Y, Iwamoto M, et al. (1999). Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic acids research. 27: 297-300.
- Kiran K, Ansari SA, Srivastava R, et al. (2006). The TATA-box sequence in the basal promoter contributes to determining light-dependent gene expression in plants. Plant Physiol. 142:364-376.
- Song J, Wang Z.(2009). Molecular cloning, expression and characterization of a phenylalanine ammonia-lyase gene (SmPAL1) from Salvia miltiorrhiza. Mol. Biol. Rep.36:939-952.
- Donald RG,Cashmore AR. (1990). Mutation of either G box or I box sequences profoundly affects expression from the Arabidopsis rbcS-1A promoter. EMBO J.9:1717-1726.
- Menkens AE, Schindler U, Cashmore AR.(1995). The G-box: A ubiquitous regulatory DNA element in plants bound by the GBF family of bZIP proteins. Trends Biochem. Sci.20:506-510.
- Van Herpen TW, Goryunova SV, van der Schoot J, et al. (2006). Alpha-gliadin genes from the A, B, and D genomes of wheat contain different sets of celiac disease epitopes. BMC genomics.7.
- Guillaumie S, Charmet G, Linossier L. (2004). Colocation between a gene encoding the bZip factor SPA and an eQTL for a high-molecular-weight glutenin subunit in wheat (Triticumaestivum). Genome.47:705-713.

Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences